-
转座子(transposable elements,TEs)被定义为能够在生物体基因组中移动的DNA序列,能在同一染色体的不同位点或者不同染色体之间转移[1]。由于起源和进化路径的差异,TEs包含不同的家族。FINNEGAN[2]首次根据TEs的转座中间体和转座机制将转座子分为Ⅰ类RNA转座子(retrotransposons)和Ⅱ类DNA转座子(DNA transposons)。Ⅰ类通过RNA介导的复制-粘贴过程迅速增殖,RNA转座子进一步分为:长末端重复序列反转录转座子(long terminal repeat,LTR,也称为内源性逆转录病毒)、非LTR反转录转座子(non-LTR)、PLEs(penelope-like elements)、DIRS(dictyostelium intermediate repeat sequence)[3]。Ⅱ类使用剪切-粘贴机制增加拷贝数[4-5],包括末端反向重复序列(terminal inverted repeat,TIR)、微型反向重复序列转座子(miniature inverted repeat transposable elements,MITEs)和Helitrons[2]。自然选择和遗传漂变导致TEs在不同物种中类别的比例和含量都不相同,在同一物种的个体之间也存在差异[6]。研究表明:人类基因组大约一半为TEs[7],其中RNA反转座子约42%[8],LTR反转座子约8%[9];在小鼠Mus musculus和人类的基因组中,长散在核元件(long interspersed nuclear elements-1,LINE-1)大约20%[10];小麦Triticum aestivum和小麦白粉病真菌Blumeria graminis基因组中,90%的序列是TEs[11];水稻Oryza sativa转座子的20%~40%中,Ⅱ类DNA转座子含量甚至高于Ⅰ类RNA转座子4倍以上[12],其中LTR约14%,而non-LTR反转座子却只有1%[13]。在玉米Zea mays基因组中,TEs含量高达85%,其中LTR反转座子和其他TEs家族含量分别为70%和15%[14-15]。通常,TEs对宿主有很多积极的影响。例如,TEs的插入控制着包括牵牛花Ipomoea purpurea在内的所有花色变化[16],贡献了可供选择的性状。反转录转座子的正常转座不仅可以产生果肉呈红色的血橙Citrus sinensis[17],还控制着葡萄Vitis vinifera[18]和番茄Solanum lycopersicum[19]等果实的颜色和形状,也参与着番茄茎尖分生组织的形成[20],还影响着哺乳动物骨骼的发育[21]。并且,可以利用TEs的激活诱导疾病的发生,从而明确疾病的机理,寻找出治疗的药物与方法。然而,由于TEs的负面影响而被称为“垃圾DNA”。例如,LINE-1是人类基因组中唯一的自主转座元件,它的表达成为许多恶性肿瘤的标志[22],并且导致包括精神分裂症在内的众多精神疾病[23],人类的120多种遗传疾病都是由于LINE-1的插入而引起的[24],其拷贝数的增加会导致腺瘤性息肉病基因(APC)肿瘤抑制基因突变从而引发人类直肠癌(colorectal cancer,CRC)[25]。ZmNAC111
基因是维持玉米幼苗耐旱性的关键基因,MITE转座子的插入会下调ZmNAC111的表达,从而引起玉米幼苗的干旱敏感性增强[26]。在小鼠生殖系中,TEs增加拷贝数会导致其不育[27],并有调控具有双向命运细胞的潜能[28]。TEs插入基因组中不仅破坏基因的功能,而且对邻近基因的表达有极性影响[29],对着丝粒稳定性同样具有重要的作用。由此可见,TEs转座破坏了宿主基因组的稳定,也搅动了宿主的基因表达调控网络,因此,TEs活性通常受到宿主多种表观遗传修饰机制的调控,例如,DNA甲基化、抑制性组蛋白修饰、小RNA途径和染色质途径。DNA甲基化是高等真核生物中广泛存在的保持TEs沉默的表观遗传修饰方式,包括从头甲基化、维持甲基化和脱甲基3个水平[30]。哺乳动物基因组中主要为CG二核苷酸序列环境的胞嘧啶甲基化,由DNA甲基转移酶1(DNA methyltransferase 1,DNMT1)和DNA甲基转移酶3(DNA methyltransferase 3,DNMT3)维持,植物中还具有CHG和CHH(H表示A、T或C)胞嘧啶环境的甲基化[31],则是由与DNMT3相似的域重排甲基转移酶1(domains rearranged methyltransferase 1,DRM1)和域重排甲基转移酶2(domains rearranged methyltransferase 2,DRM2)催化[32]。本研究论述了TEs沉默与DNA甲基化的关系,重点总结了以DNA甲基化为主的转座子沉默机制最新研究进展,归纳了环境因素通过DNA去甲基化调控转座子跳跃的机理。
HTML
-
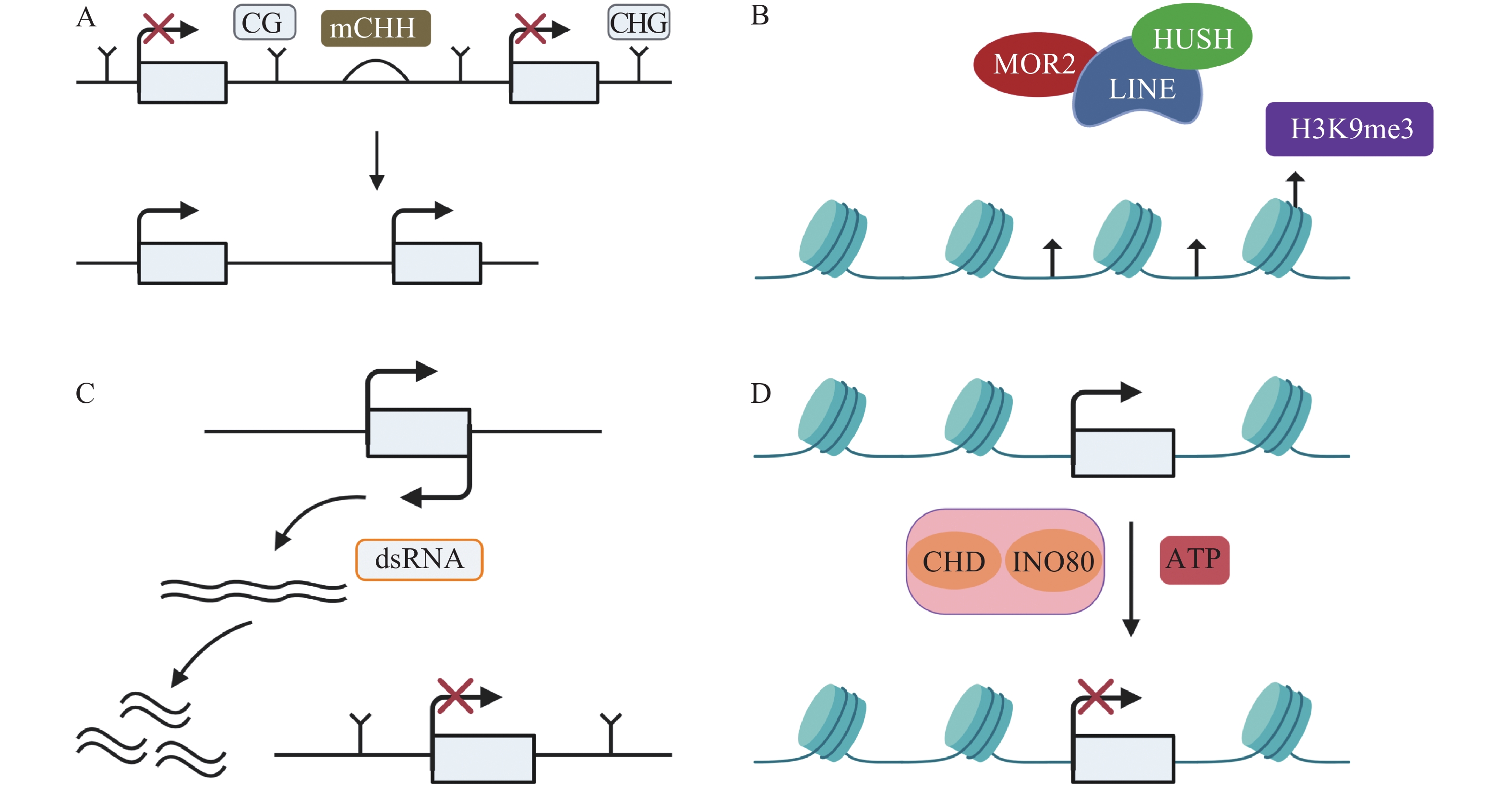
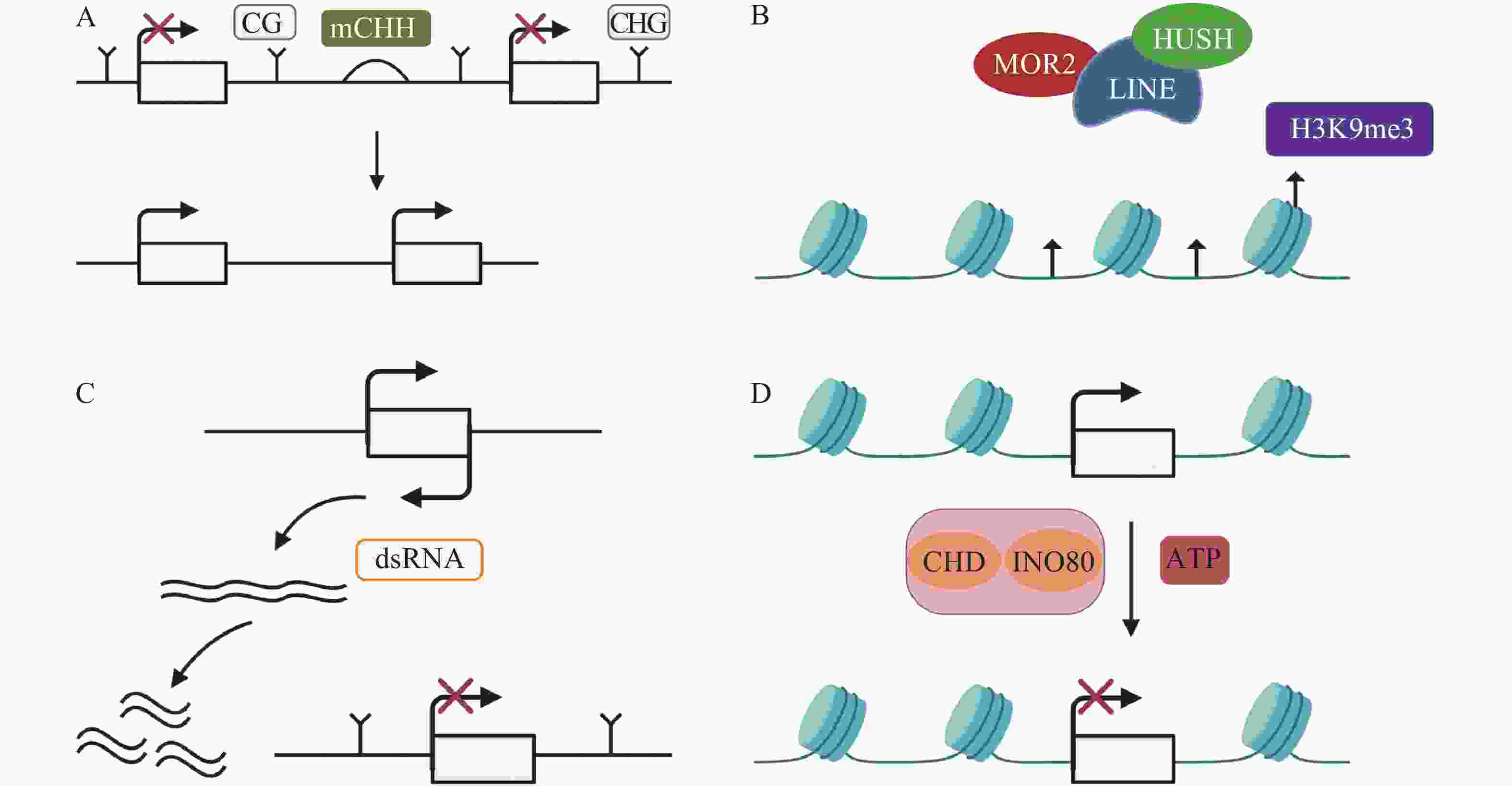
TEs沉默分为检测、扩增和抑制3个部分[33],保持TEs沉默通常受DNA甲基化、抑制性组蛋白修饰、小RNA途径以及染色质途径的调控。(1)例如,在玉米基因组中,mCHH甲基化岛常常插入活跃基因与沉默转座子之间,去甲基化会导致沉默的转座子表达上调,RNA指导的DNA甲基化(RNA-directed DNA methylation,RdDM)能够维持转座子的沉默[34],mCHH甲基化岛缺失会导致CG、CHG的丢失,同时上调TEs活性(图1A)。DNMT1在hNPCs (human neural progenitor cells)维持DNA甲基化,通过CRISPR-Cas9技术去除DNMT1后,导致CPG甲基化水平降低,激活LINE-1,进一步影响与精神疾病有关的基因[35]。(2)抑制性组蛋白修饰是另一个沉默TEs的途径。通常认为组蛋白H3的赖氨酸9和27的三甲基化H3K9me3 (histone H3 Lys9 trimethylation)、H3K27me3 (histone H3 Lys27 trimethylation)能够沉默TEs。MORC2蛋白和HUSH (human silencing hub)与在进化上较年轻的全长LINE-1结合,诱导组蛋白H3K9me3富集,从而沉默TEs(图1B)[36]。水稻的H3K4特异性脱甲基酶蛋白JMJ703介导H3K4脱甲基,当JMJ703活性受到影响时,增加H3K4me3积累,2个LINE元素被激活转座[37]。(3)小RNA途径同样是沉默TEs的有效途径。AT(alternative transposition)产生的CIs(composite insertions)的反向复制被转录生成dsRNA(double-stranded RNA),等位基因P1-WW-ID1和P1-WW-ID4上富集21、22、24 nt (nucleotide) siRNA,然后siRNA介导玉米Ac/Ds转座子沉默(图1C)。这是首次提出TEs自主介导的沉默[38]。RNA与Piwi (P-element-induced wimpy testis)蛋白相互作用结合形成piRNAs,Hsp70伴侣蛋白是piRNAs生物合成的主要参与者,在果蝇Drosophila生物体中,Hsp70伴侣蛋白遭受热激胁迫导致piRNAs的合成被破坏,因此在转录后水平增加了TEs的表达[39]。(4)染色质途径对TEs的沉默同样也很重要。染色质重塑复合物(chromatin remodelers)包括CHD、SWI/SNF、INO80、SWR1等,它能利用ATP水解的能量移动或者重组核小体,从而沉默TEs(图1D)[40-41]。SWI/SNF家族中SWI3B协同HDA6 (histone deacetylase 6)增加H3K9me2水平,沉默TEs,同时,MET1和SUVH4/5/6也参与增加H3K9me2以及DNA甲基化维持TEs沉默[42]。
-
DNA甲基化在TEs沉默中的作用已被很多研究证实。毛竹Phyllostachys edulis的DNA甲基化水平经过甲基化抑制剂5-氮杂胞苷和γ射线的处理后显著降低,具有转座活性的MITEs家族转座子PhTst-3-79的转座频率相比野生型对照显著增加,并且DNA甲基化随甲基化抑制剂浓度和γ射线辐照剂量增加而下降,TEs的转座频率也随之增加[43]。ZHOU等[44]鉴定的毛竹基因组全长LTR反转录转座子PHRE2(Phyllostachys edulis retrotransposon 2)经过脱落酸(ABA)、水杨酸(SA)、γ射线处理后,甲基化水平显著降低,而拷贝数显著增加。
转录后基因沉默(post-transcriptional gene silencing,PTGS)以RDR6合成的双链RNA为起始,然后在由DCL2/4(dicer-like 2/4)介导产生21~22 nt (nucleotide) sRNA(图2A),招募DRM1/2产生5-甲基胞嘧啶(5mC)[45]。其中,21 nt sRNA在转录后水平和24 nt sRNA在转录水平指导TEs沉默[46]。RdDM是开花植物维持TEs沉默的主要机制[47]。RdDM是由RNA聚合酶Ⅳ (RNA polymerase Ⅳ,Pol Ⅳ)介导RNA转录起始,依赖RNA的RNA聚合酶2 (RDR2)合成双链RNA,然后双链RNA在RDR2和DCL3(dicer-like 3)作用下,降解为24 nt sRNA,AGO4/6 (ARGONAUTE 4/6)蛋白与24 nt sRNA结合,最终由DRM1/2介导DNA甲基化(图2B)[31, 48-49]。最新的关于植物non-CG甲基化的研究中,在转座子失活的3个阶段基础上,提出了第4个阶段[45]。在番茄基因组中,第1阶段为转录后基因沉默,LTR反转座子在Pol II(RNA polymerase II)等参与下,生成21~22 nt sRNA(small RNA)(图2A)。第2阶段是RdDM的短暂参与,LTR拷贝数增加,RdDM导向LTR甲基化(图2B)。第3阶段不包含RdDM途径,而是由MET1和CMT3维持沉默(图2C)[50]。第4阶段中,沉默的LTR反转座子失去转座能力,不再受MET1和CMT3的靶向,再次开始转录,RdDM第2次增加LTR反转座子甲基化水平(图2D)。
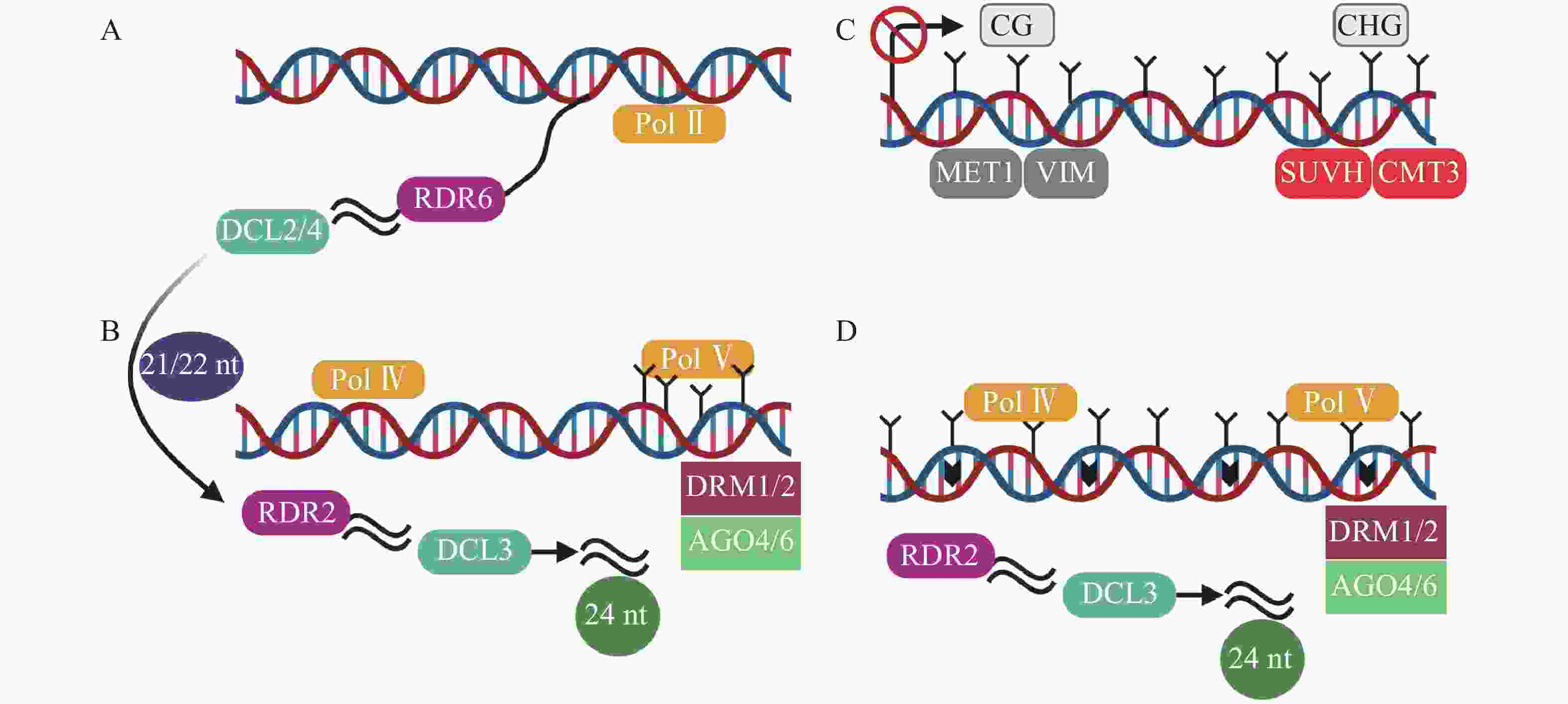
Figure 2. Four stages of TEs silence in tomato[45]
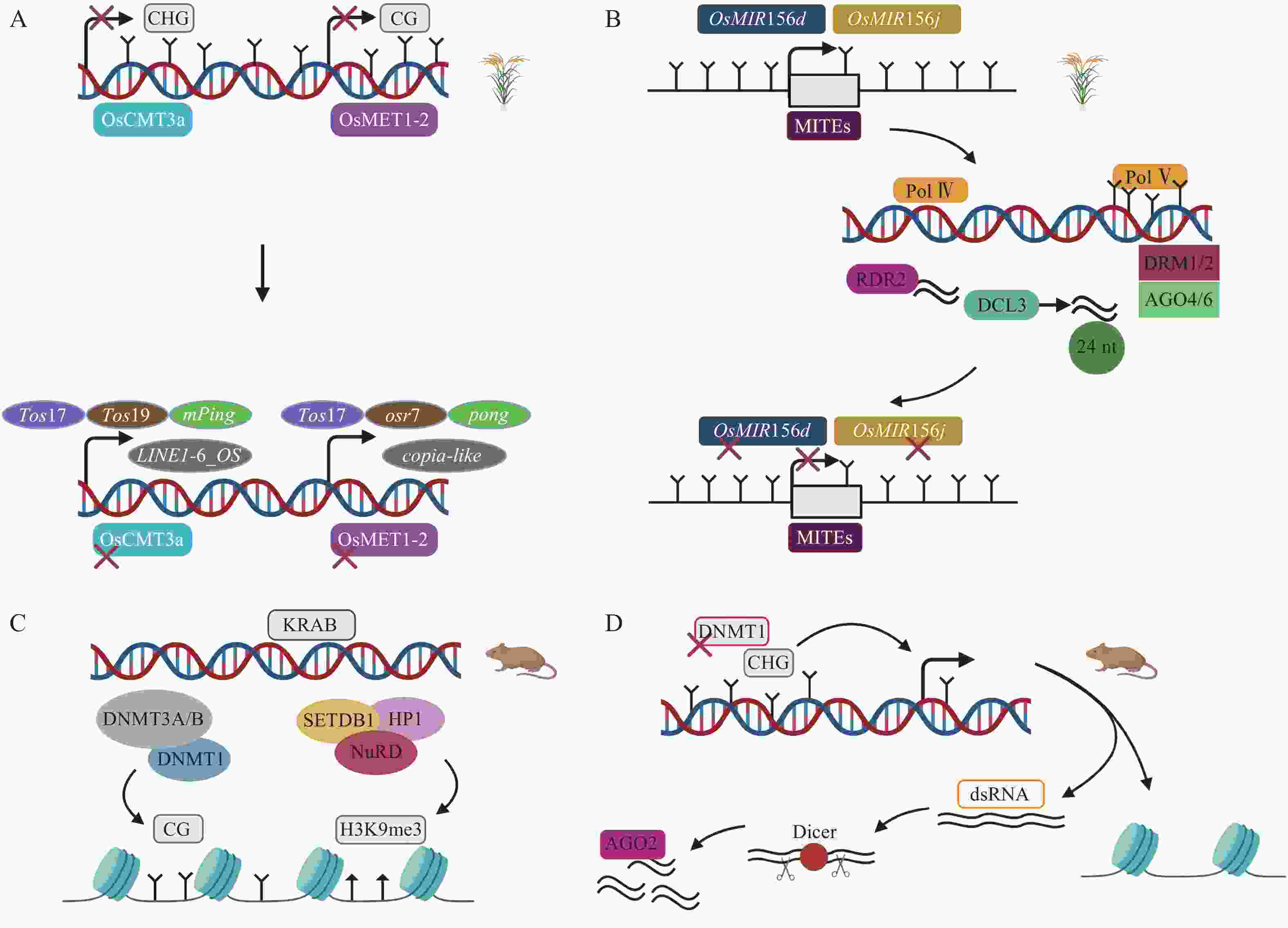
甲基化酶对DNA甲基化的维持是非常重要的,间接调控TEs的活性。在水稻基因组中,染色体甲基化酶OsCMT3a维持CHG甲基化维持TEs沉默,转座子Tos17处理的OsCMT3a突变体甲基化水平降低,繁殖阶段时,8个TEs家族发生转座,其中包括1个LINE,1个MITE等[51]。关于水稻基因组甲基化水平下调而沉默TEs的研究中,在DNA甲基转移酶OsMET1-2纯合突变体中发现,CG甲基化损失激活包括低拷贝LTR反转座子copia-like在内的TEs[52](图3A)。在拟南芥Arabidopsis thaliana中,关于RdDM通路的研究已有很多,但很少有明显RdDM通路介导的发育表型变化。RdDM途径影响水稻TEs表达从而导致水稻表型发生变化。OsMIR156d和OsMIR156j是水稻中促进分蘖的基因,其启动子区域的MITEs被RdDM介导甲基化,抑制OsMIR156d和OsMIR156j基因的表达,从而调控水稻的分蘖[49](图3B)。这在表观遗传水平上对调控农艺性状的表达具有重要意义。另一个沉默TEs的关键通路涉及KRAB-ZFPs(krüppel-associated box-zinc-finger proteins)。在小鼠胚胎发生早期,KRAB-ZFPs特异识别TEs,KAP1(KRAB-associated protein 1)作为辅助因子,在组蛋白甲基转移酶SETDB1(SET domain bifurcated 1)、HP1(heterochromatin protein 1)以及组蛋白去乙酰化酶复合物NuRD(nucleosome remodeling deacetylase)等作用下形成压制性染色质结构,维持H3K9me3,抑制内源性逆转录病毒(endogenous retrovirus,ERVs),胚胎干细胞(embryonic stem cells,ESCs)KAP1缺失将导致ERVs的上调,并且,DNMT1、DNMT3A/B也参与沉默ERVs,但是敲除DNMT1、DNMT3a/b后,KRAB-ZFPs仍然维持绝大多数ERVs的沉默[53-55](图3C)。小RNA途径可以作为TEs激活后的快速防御,而抑制性组蛋白修饰作为接下来的缓慢防御。小鼠ESCs中,DNMT1缺失导致CPG甲基化水平从85%降到20%,DNA去甲基化诱导TEs激活,这是因为低甲基化时的反义TEs转录,核酸内切酶Dicer切割dsRNA,接下来AGO2(ARGONAUTE2)与小RNA结合,基于endosiRNA(endogenous short interfering RNAs)的抑制机制沉默甲基化丧失激活的TEs[56](图3D)。
无论Ⅰ类或Ⅱ类TEs的活性,都与DNA甲基化水平密切相关。En/Spm DNA家族转座子也称CACTA转座子[57]。在红肉萝卜Raphanus sativus中,CACTA转座子高度甲基化导致其拷贝数下降,同时甲基化扩散至花青素合成基因RsMYB1启动子区域,导致基因RsMYB1表达下调,影响花青素的积累[58]。有研究通过分析癌症数据库,发现400多个TEs表达上调,其中包括HERVs (human endogenous retroviral)、LINE、SINE等,接近2/3的TEs表达上调似乎是由于邻近区域DNA甲基化的损失导致的[59]。敲除番茄基因组中对CHG甲基化起关键作用的KYP和CMT3基因后,LTR反转座子富集在上调基因的启动子区域[45]。DNA糖基化酶介导的主动去甲基化同样可以激活水稻反转座子[60]。反转录转座子MRL (multiretrotransposon-like)插入大麦Hordeum vulgare基因组启动子区域,可以极大增强HvAACT1基因的表达,从而增强大麦抵抗铝毒害的能力。但是MRL的插入通常伴随着高甲基化,因此只有MRL转座子去甲基化才能增强大麦耐铝性[61]。水稻中,LTR反转座子的插入导致有害的异位重组,高度甲基化抑制LTR反转座子的活性,从而抑制这种有害作用[62]。在小鼠中,胞嘧啶甲基化在缺乏DNMT1的胚胎中下调,导致内源性逆转录病毒ERVs(endogenous retrovirus)上调,这是首次证明DNA甲基化在小鼠中沉默TEs的研究[63]。丝状真菌Neurospora crassa中的Ⅰ类TEs在胞嘧啶甲基化信号诱导下发生甲基化,下调了TEs的表达[64]。在斑马鱼Danio rerio基因组中,DNA低甲基化上调RNA转座子表达[65]。
-
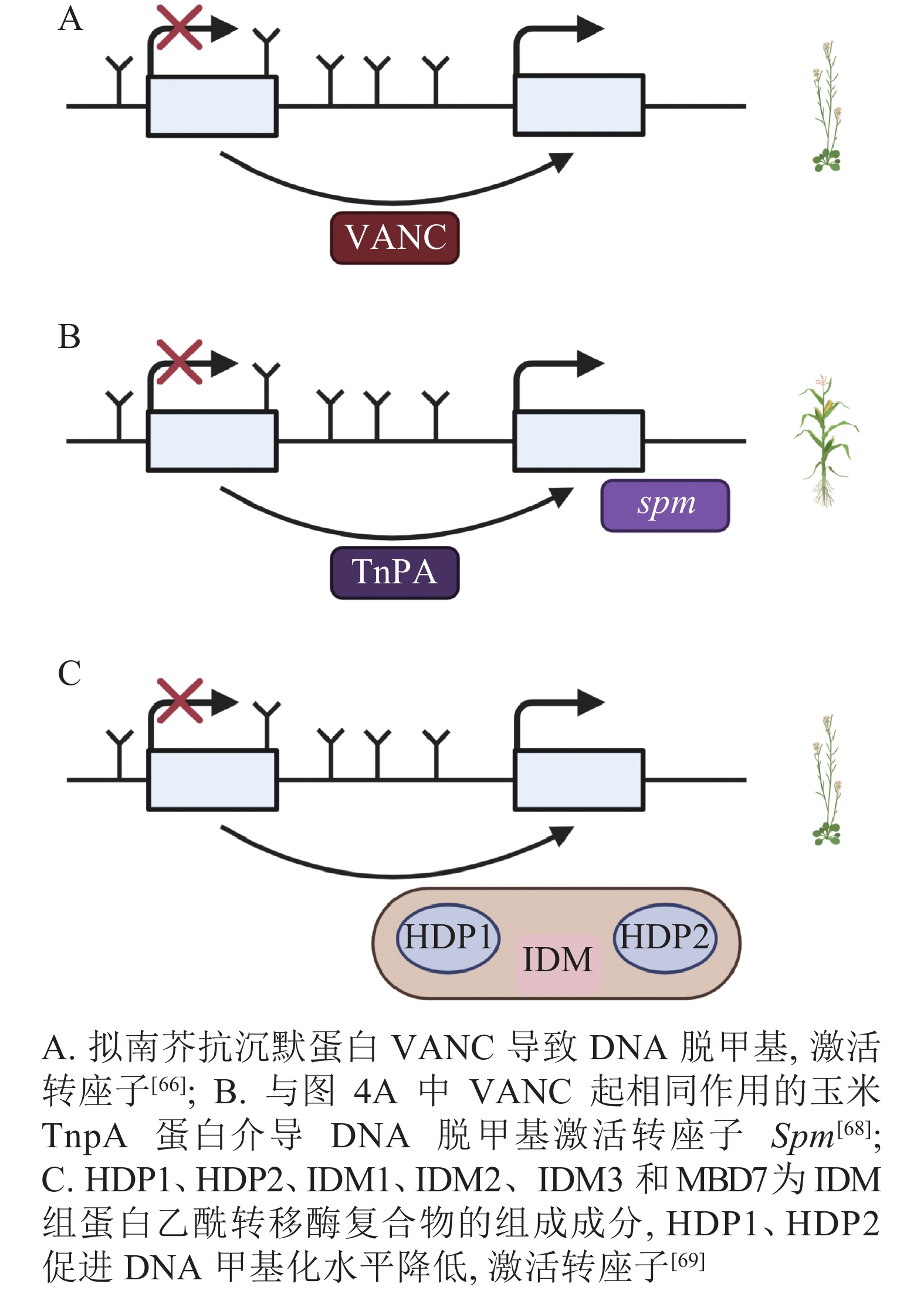
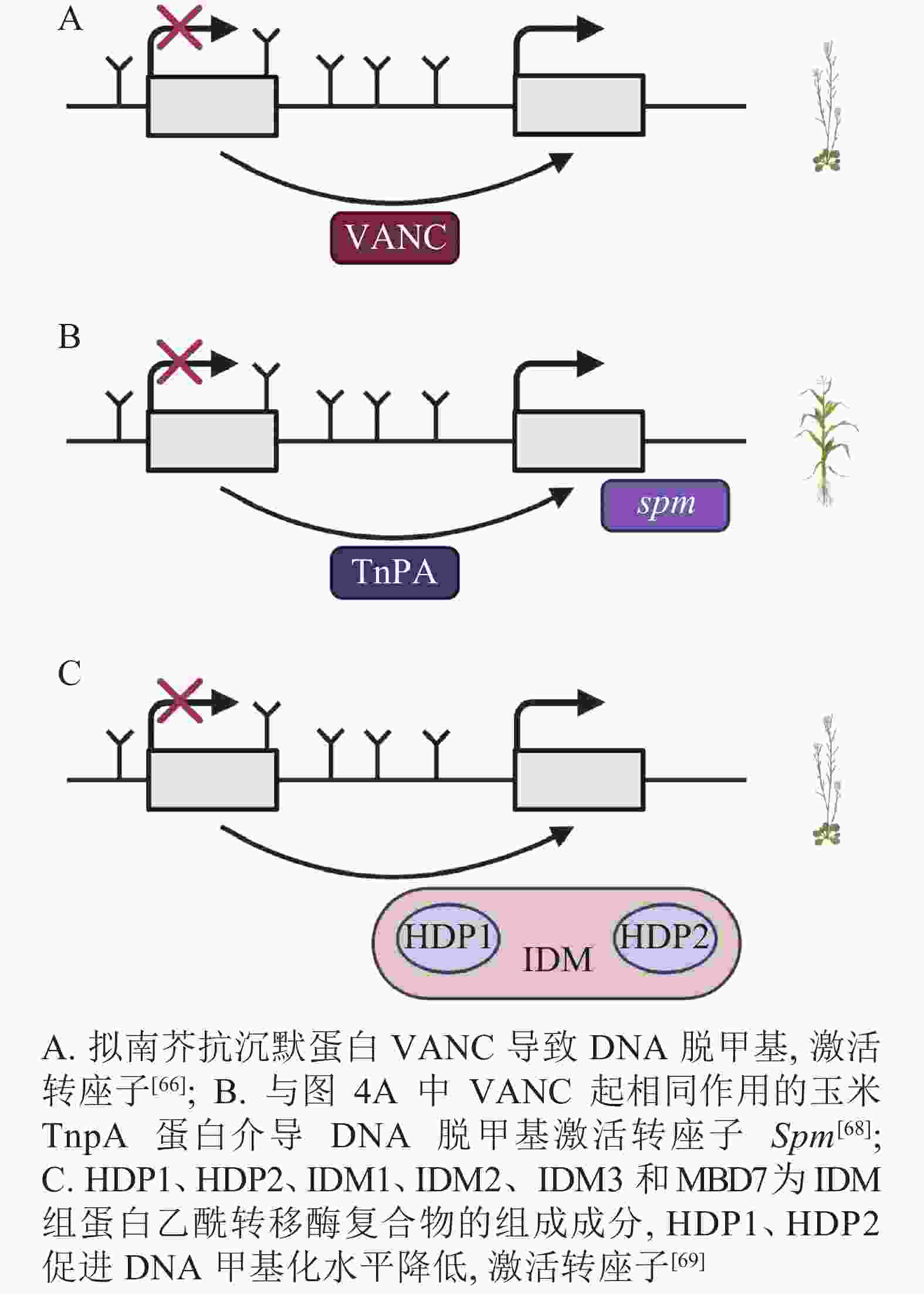
DNA甲基化是当TEs对宿主产生有害影响时的防御机制,但在拟南芥中,进化出转座子Hi (Hiun)编码的抗沉默蛋白VANC,上调被DNA甲基化沉默的TEs表达。这种抗沉默蛋白VANC不仅可以诱导低甲基化,增加TEs的拷贝数,而且能够把对宿主的不利影响降到最小[66-67](图4A)。与抗沉默蛋白VANC一样诱导低甲基化的还有玉米转座子编码蛋白TnpA,TnpA介导玉米转座子Spm DNA脱甲基化,这是由TnpA结合到Spm上,在脱甲基底物和脱甲基酶的参与下进行的DNA去甲基化[68](图4B)。在拟南芥中,Harbinger转座子编码的2个蛋白HDP1(H arbinger-derived protein 1)和HDP2(H arbinger-derived protein 2)的正常表达可以维持低甲基化和内源TEs的沉默,HDP1、HDP2、IDM1(increased DNA methylation 1)、IDM2等作为IDM(increased DNA methylation)组蛋白乙酰转移酶复合物的组成成分,任一结合因子的突变都会升高甲基化水平和影响TEs的表达。HDP1、HDP2突变后下调了AT1TE46455、AT1TE36115和AT1TE35325的表达[69-70](图4C)。
-
生物或非生物胁迫会导致DNA甲基化发生改变,例如,强烈的锌缺乏会导致拟南芥全基因组的DNA甲基化水平发生变化[71]。在CG对称环境中,缺磷对甲基化水平影响较小,但缺氮会导致玉米根甲基化水平降低[72]。水稻在遭受盐胁迫时,盐敏感的IR29缺乏改变DNA甲基化水平的能力,而耐盐水稻能降低甲基化水平[73]。双生病毒干扰植物DNA甲基转移酶MET1和CMT3,导致DNA去甲基化[74]。连作胁迫导致大豆Glycine max基因组DNA去甲基化酶ROS1和DML增加表达,从而降低基因组DNA甲基化水平[75]。
环境胁迫会降低DNA甲基化水平,可能诱导TEs激活。用鞭毛蛋白衍生肽flg22(flagellin peptide 22)处理拟南芥叶片,会触发DNA去甲基化,导致某些TEs转录增加[76]。在棉花Gossypium hirsutum基因组中,高温胁迫导致DNA甲基化水平降低,甲基化程度较高的TEs被激活增加拷贝数[77]。在拟南芥的精子伴细胞(vegetative cell,VC)中,H1组蛋白缺失会使转录起始位点发生去甲基化,从而激活TEs[78]。用去甲基化剂5-azaC处理水稻种子后,激活Dart1-24 DNA转座子,同时验证了转座子拷贝数增加是5′区域的核苷酸去甲基化导致的[79]。在磷酸盐缺乏的环境中,水稻基因组DNA甲基化水平优先在TEs中瞬时变化,在抑制TEs方面发挥潜在的作用。磷酸盐不足的压力环境诱导高甲基化,从而沉默TEs[80]。
-
通常,TEs对宿主不利影响是由于TEs的插入破坏启动子区域或基因区域,导致基因组重排,以及引起的缺失、重复、倒位等基因组结构变异[81-82]。调控TEs表达对维持基因组稳定性具有重要意义,DNA修饰是物种进化过程中普遍采用的调控TEs表达的方式。现阶段,基因组学和表观基因组学的快速发展推动了DNA修饰的研究,其中,DNA甲基化是最主要的调控机制之一。然而,DNA修饰的这种作用具有不稳定性,包括在亲缘关系很近的物种中也存在变异[31]。并且,DNA甲基化水平变化涉及特定酶的参与,影响甲基化酶发挥作用的因素也在间接影响TEs表达。因此,与DNA甲基化相关的酶和基因的作用被进一步发现,可以明确DNA甲基化调控TEs表达的机制。生物或非生物胁迫导致的甲基化水平降低以及主动去甲基化上调TEs,TEs抵抗由DNA甲基化介导的沉默,编码了例如VANC、TnpA等抗沉默蛋白,促进DNA脱甲基从而增加TEs拷贝数[67-68]。宿主在遗传进化过程中更好适应环境的前提就是基因组的稳定,TEs的表达则是破坏基因组稳定的重要因素,环境因子诱导DNA甲基化水平发生变化,进一步影响TEs的表达。DNA甲基化和TEs响应环境因子的互作机制,以及最终调控宿主基因表达的机制是未来研究的方向。









 DownLoad:
DownLoad: