-
小麦Triticum aestivum作为全球三大粮食作物之一,其种植面积约占全球可耕地面积的22%,为超过35%的世界人口提供主食来源,对保障全球粮食安全具有战略意义[1]。抽穗期是小麦从营养生长转变到生殖生长的关键转折点,其适时性直接影响植株的光能利用效率、籽粒灌浆进程及最终产量形成,同时决定作物对特定生态区的适应性[2]。解析抽穗期的遗传调控机制,不仅可为不同地区制定最佳播期提供理论依据,通过培育早熟品种实现与水稻Oryza sativa、玉米Zea mays等作物的合理轮作,优化土地资源配置,更对提升种植效益和农业可持续发展具有重要意义[3]。
作为复杂数量性状,小麦抽穗期主要受光周期[4]、春化[5]、自身早熟性[6]三大遗传通路的协同调控,已有研究通过连锁作图与关联分析,在小麦各同源群染色体上定位到了多个控制抽穗期的主效数量性状位点(QTL),并通过功能基因组学手段验证了关键调控因子的生物学功能[7−9]。六倍体普通小麦(2n=6x=42,AABBDD),在其进化过程中发生了2次异源多倍化事件[10],其四倍体祖先野生二粒小麦Triticum turgidum var. dicoccoides (2n=4x=28,AABB)因具有抗旱、抗病、耐盐碱、品质高等优良性状,是小麦遗传改良的“天然基因库”[11]。之前诸多研究都对野生二粒小麦的相关基因进行了挖掘和功能验证,并用于小麦遗传改良[12−15]。以色列魏茨曼科学研究所FELDMAN教授团队利用染色体端体介导的同源重组技术和细胞学鉴定等方式,创制了一套普通小麦-野生二粒小麦染色体臂置换系(chromosome arm substitution lines,CASLs),其中CASL3AL携带来自野生二粒小麦的3AL染色体臂片段,为‘中国春’T. turgidum ‘Chinese Spring’ (CS)遗传背景下解析外源染色体片段功能提供了独特材料[16]。
前期研究发现CASL3AL在多元种植环境条件下均表现出稳定的早熟特性[17]。为深入挖掘野生二粒小麦促进普通小麦早抽穗的候选基因,本研究从CASL3AL×CS构建的重组自交系(RILs)中筛选出2个携带3AL染色体片段的纯合早熟株系1017-7-167-1和1017-10-311-15-5,在幼穗发育的关键节点单棱期(W1.5)和二棱期(W2.5),进行转录组测序(RNA-Seq),通过单核苷酸多态性(SNP)分析精确界定外源染色体片段的导入位置,并通过基因差异表达分析筛选早抽穗相关候选基因。
-
供试材料包括CASL3AL×CS F5 (1017-7-167-1)、F6 (1017-10-311-15-5),以及普通小麦品种CS。选取颗粒饱满的种子置于发芽盒中进行发芽处理,种子露白后,将其置于4 ℃培养箱中进行20 d的春化处理。春化处理结束后,将幼苗移栽至32孔穴盘中,并置于25 ℃、长日照(16 h光照/8 h黑暗)的培养室中,让幼苗正常生长。该处理条件能够确保实验材料的生理状态一致,为后续实验提供可靠基础。
-
选取供试材料从单棱期进入二棱期阶段生长发育速度差异最大的时期,根据WADDINGTON等[18]研究的标准,对小麦幼苗的单棱期(转换顶点时期)和二棱期(小麦分生组织出现)的幼穗进行标记,分别标记为W1.5和W2.5,并在这2个时间点对供试材料的叶片进行取样。
在体视显微镜下观察幼穗生长状况,当供试材料生长至单棱期和二棱期时,分别选取同一材料3株幼苗的顶端叶片,混合后作为1个生物学重复。每个材料在单棱期和二棱期各取3个生物学重复。取样后迅速将叶片放入液氮中冷冻,以确保RNA的完整性。RNA提取、cDNA文库构建和测序工作委托武汉贝纳科技有限公司完成。
-
为确保测序数据的质量和准确性,首先利用Fastp[19]对原始数据进行质量控制,去除低质量读段和接头序列。随后,使用Hisat2[20]软件将过滤之后的高质量测序数据与CS参考基因组进行比对,将得到的Sam文件转化为Bam文件,利用Samtools[21]软件取到唯一比对,以确保测序数据的准确映射,利用featureCounts[22]对mapped reads进行表达定量。通过GATK[23]软件去除PCR重复,并对SNP变异进行检测。最终,将得到的SNP变异数据进行图表绘制,以便于后续的分析和展示。主成分分析(PCA)使用R语言中的PCAtools包,对转录组数据进行降维分析。
-
为识别单棱期和二棱期具有显著差异表达的基因,使用R语言中的DESeq2 (1.42.1)包[24]进行差异表达基因分析。在分析过程中,以对数转换的表达差异倍数(log2FC)≥1且调整后的P(Padj)≤0.05作为筛选标准,以确保筛选出的差异表达基因具有统计学意义和生物学相关性。这些筛选出的差异表达基因将用于后续的功能分析和候选基因筛选。
-
为了深入理解差异表达基因的功能,使用Triticeae-Gene-Tribe平台进行基因本体(GO)富集分析,以揭示这些基因在生物过程、分子功能和细胞组分方面的富集情况。使用R语言中的ggplot2 (3.5.1)包对GO富集分析结果进行可视化。
-
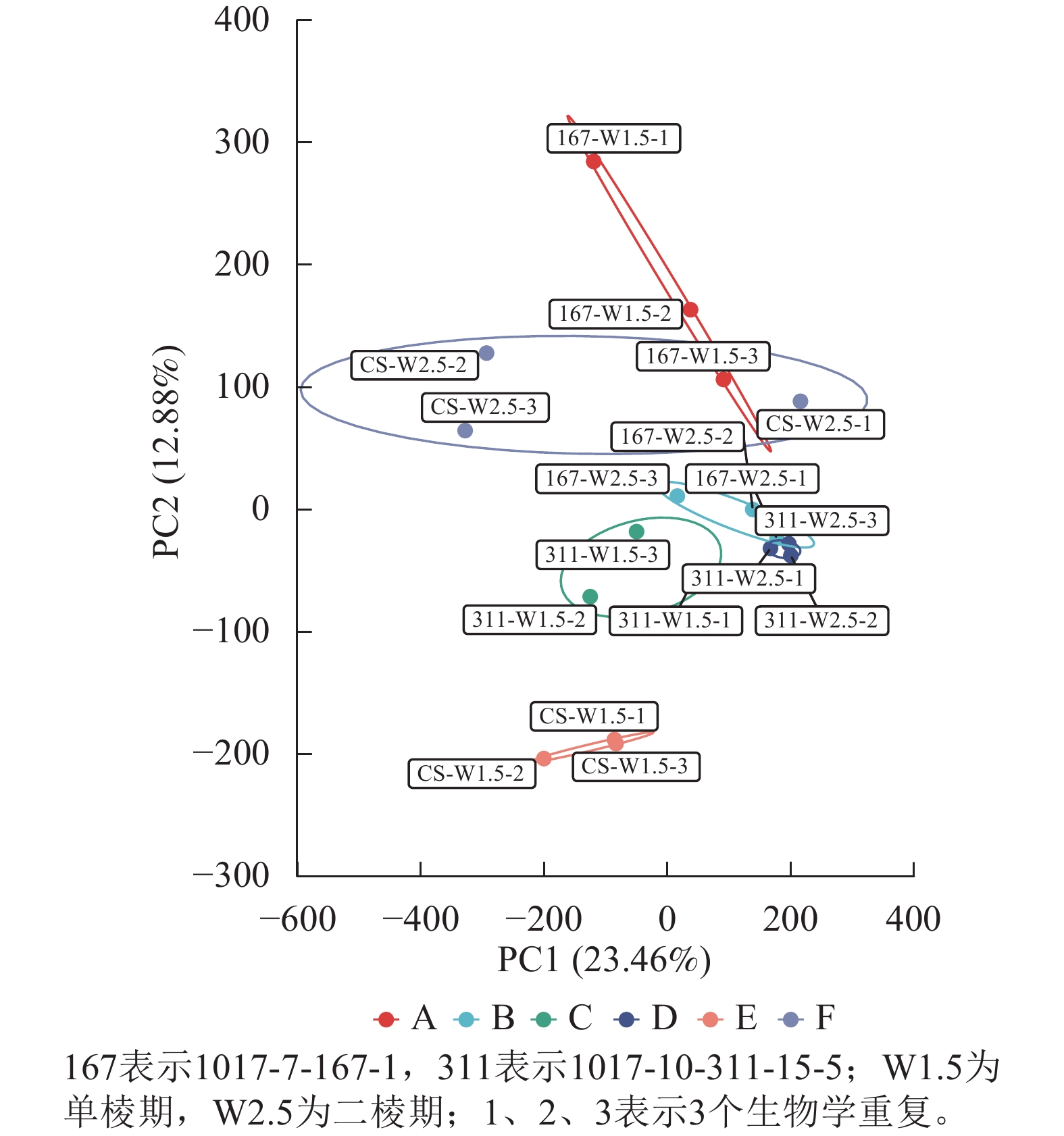
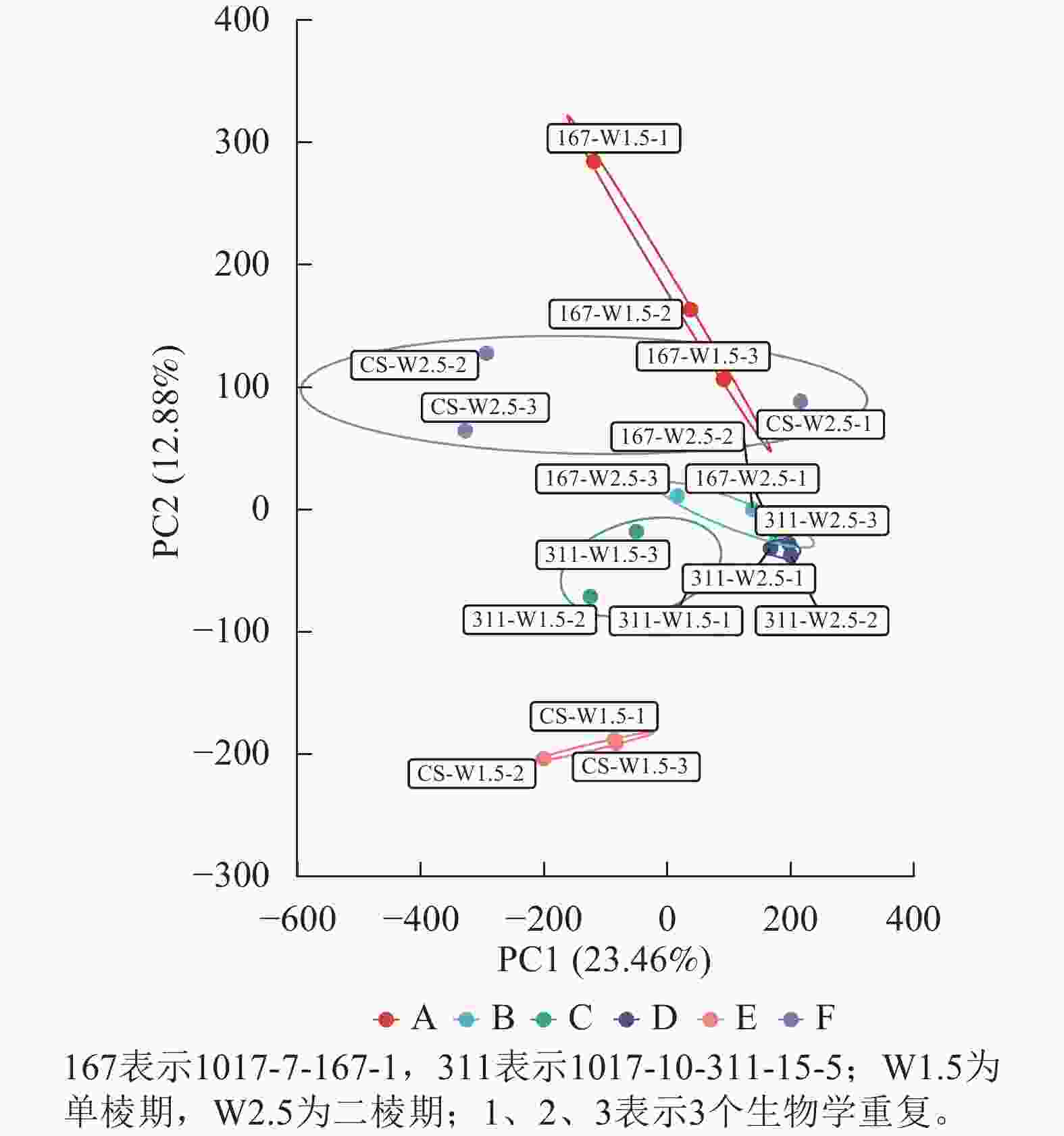
经过质量控制后,共得到260 Gb的干净数据(clean data),读长(reads)为150 bp,样品的Q30碱基百分比为94.31%~96.53%,GC含量为51.06%~53.7%,表明样品数据的可靠性高。对转录组数据进行PCA分析,结果表明同组内样品呈现聚集(图1)。
-
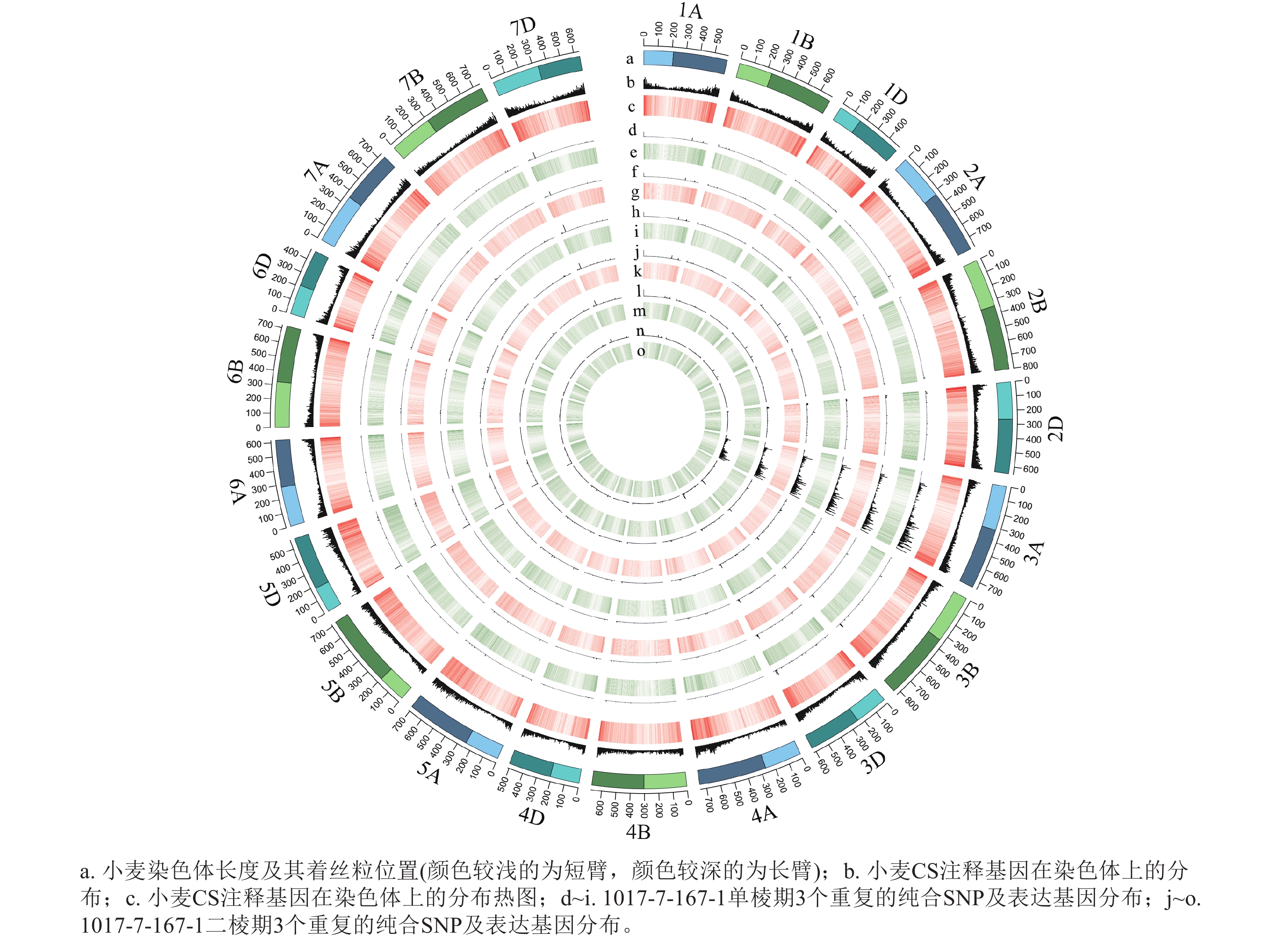
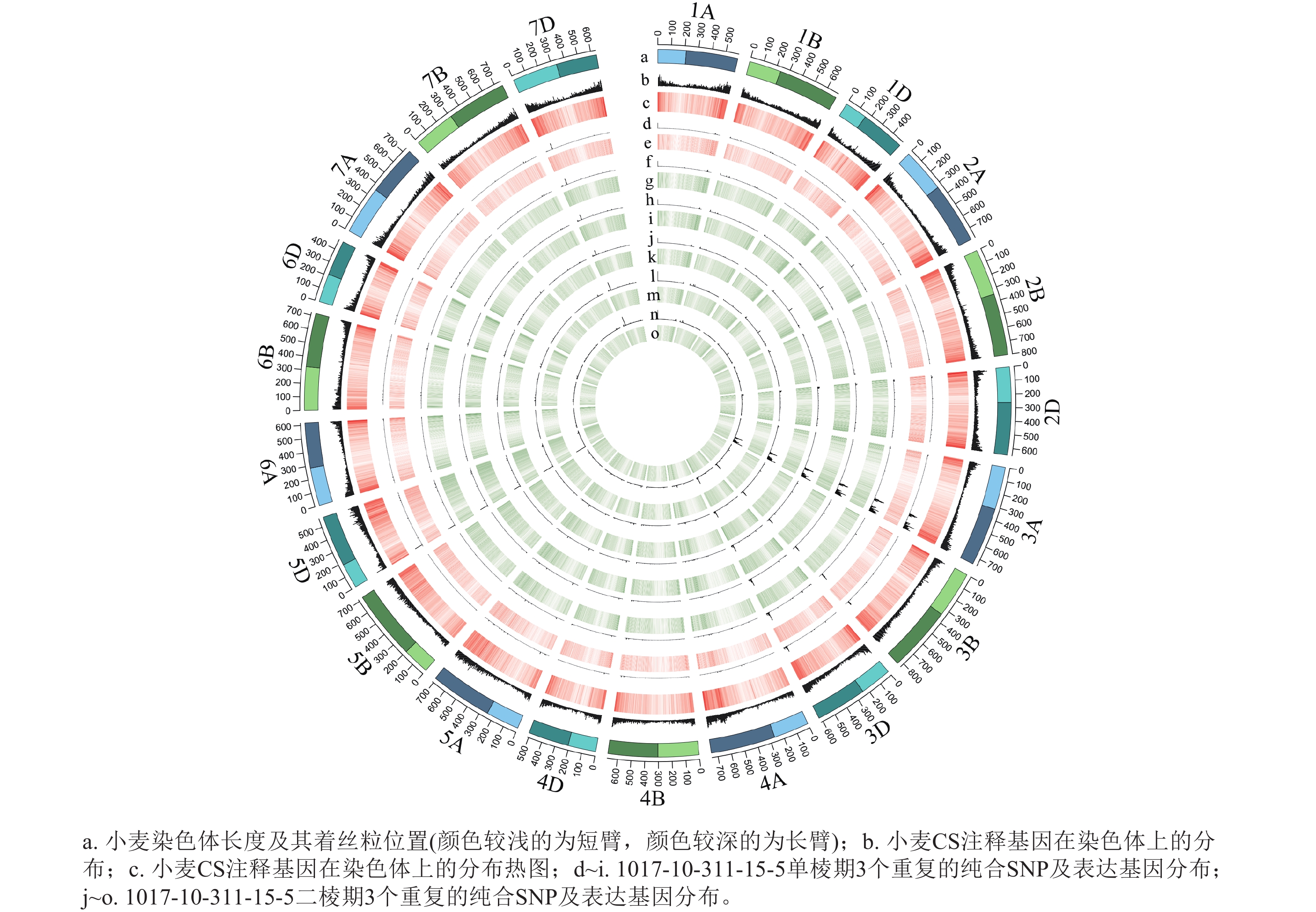
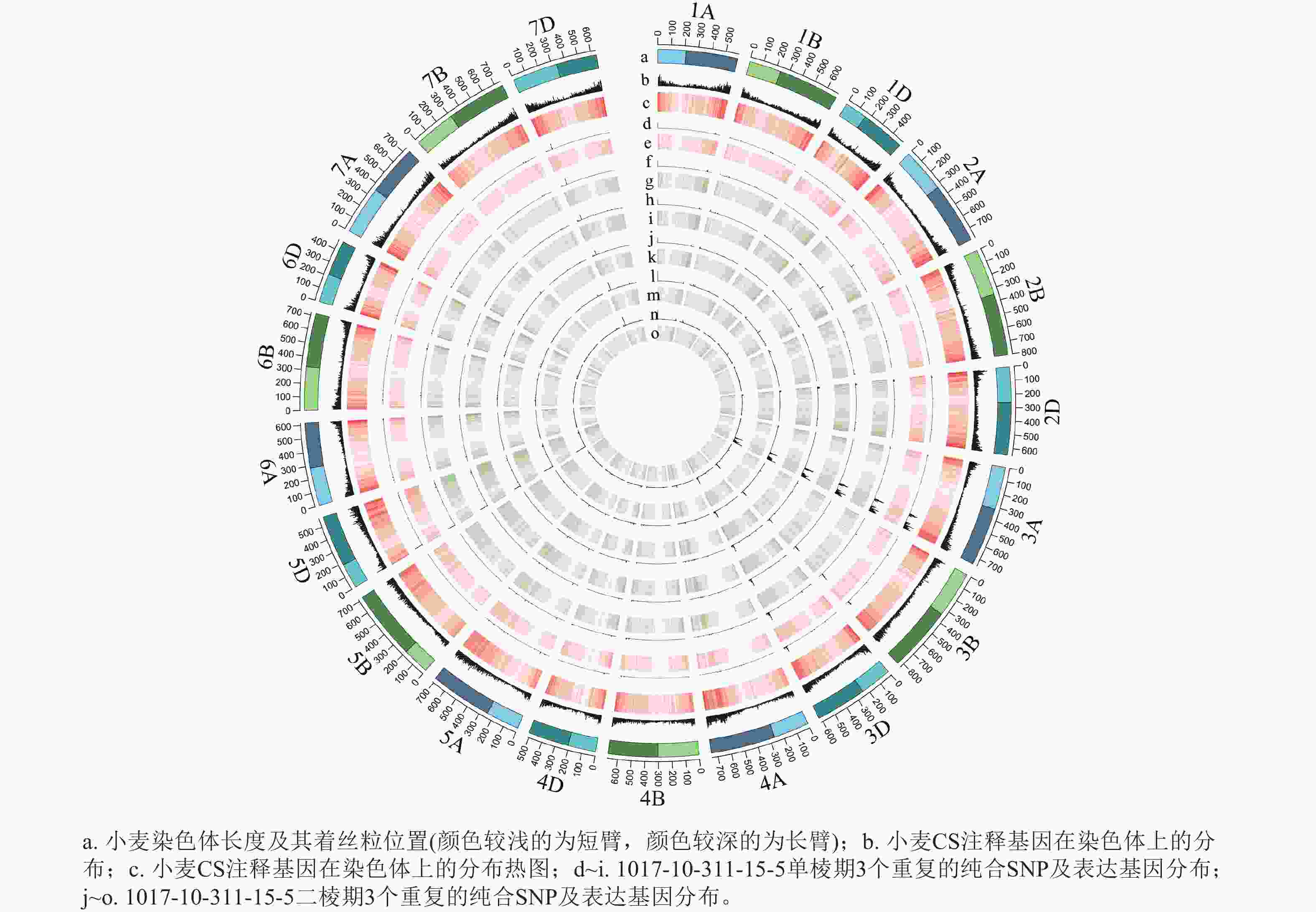
从表1可见:1017-7-167-1在单棱期共有6 497个纯合SNP,在二棱期共有4 954个纯合SNP,这些纯合SNP主要集中在3A染色体上。1017-10-311-15-5在单棱期共有3 276个纯合SNP,在二棱期共有2 829个纯合SNP,这些纯合SNP也主要位于3A染色体上。这表明在不同发育阶段,纯合SNP的分布具有一定的规律性,且主要集中在特定的染色体区域。
染色体 数量 1017-7-167-
1-W1.51017-7-167-
1-W2.51017-10-311-
15-5-W1.51017-10-311-
15-5-W2.5Chr 1A 152 126 122 132 Chr 1B 127 108 113 86 Chr 1D 121 72 79 92 Chr 2A 122 160 109 100 Chr 2B 163 123 109 101 Chr 2D 156 145 119 144 Chr 3A 3 602 2 606 898 683 Chr 3B 243 212 207 205 Chr 3D 290 171 246 187 Chr 4A 118 106 102 89 Chr 4B 81 70 79 50 Chr 4D 68 69 72 52 Chr 5A 152 110 139 80 Chr 5B 126 78 85 69 Chr 5D 203 195 179 177 Chr 6A 86 59 75 68 Chr 6B 127 110 108 80 Chr 6D 119 86 80 87 Chr 7A 149 104 125 93 Chr 7B 119 100 89 82 Chr 7D 173 144 141 172 总计 6 497 4 954 3 276 2 829 说明:Chr为染色体,W1.5为单棱期,W2.5为二棱期。 Table 1. Quantitative distribution of homozygous SNPs on chromosomes
-
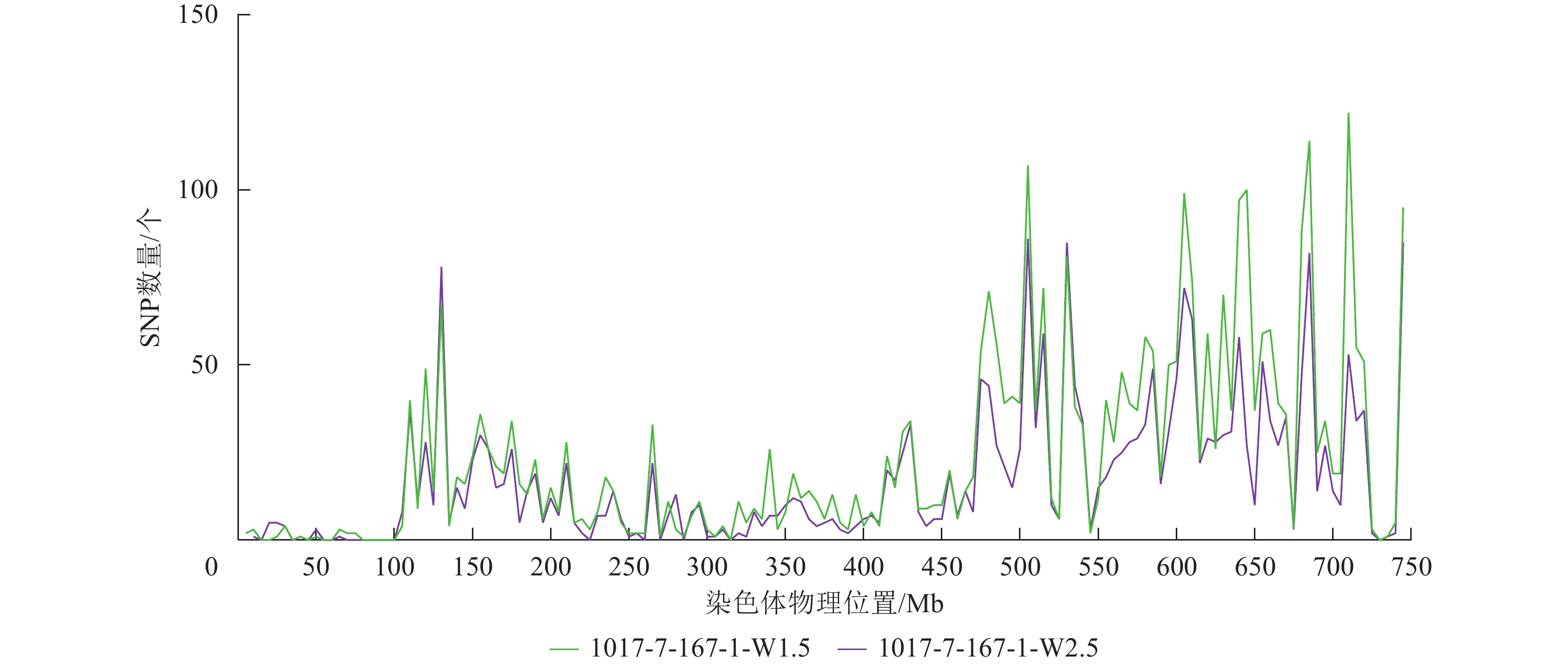
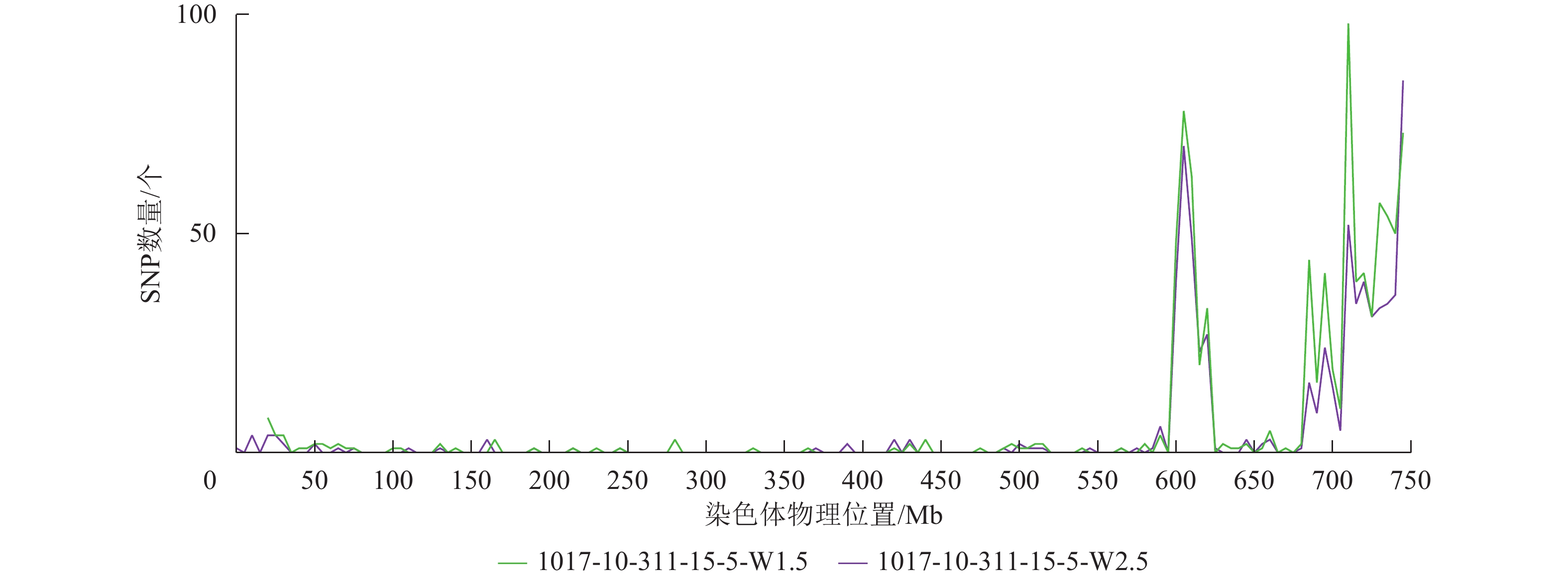
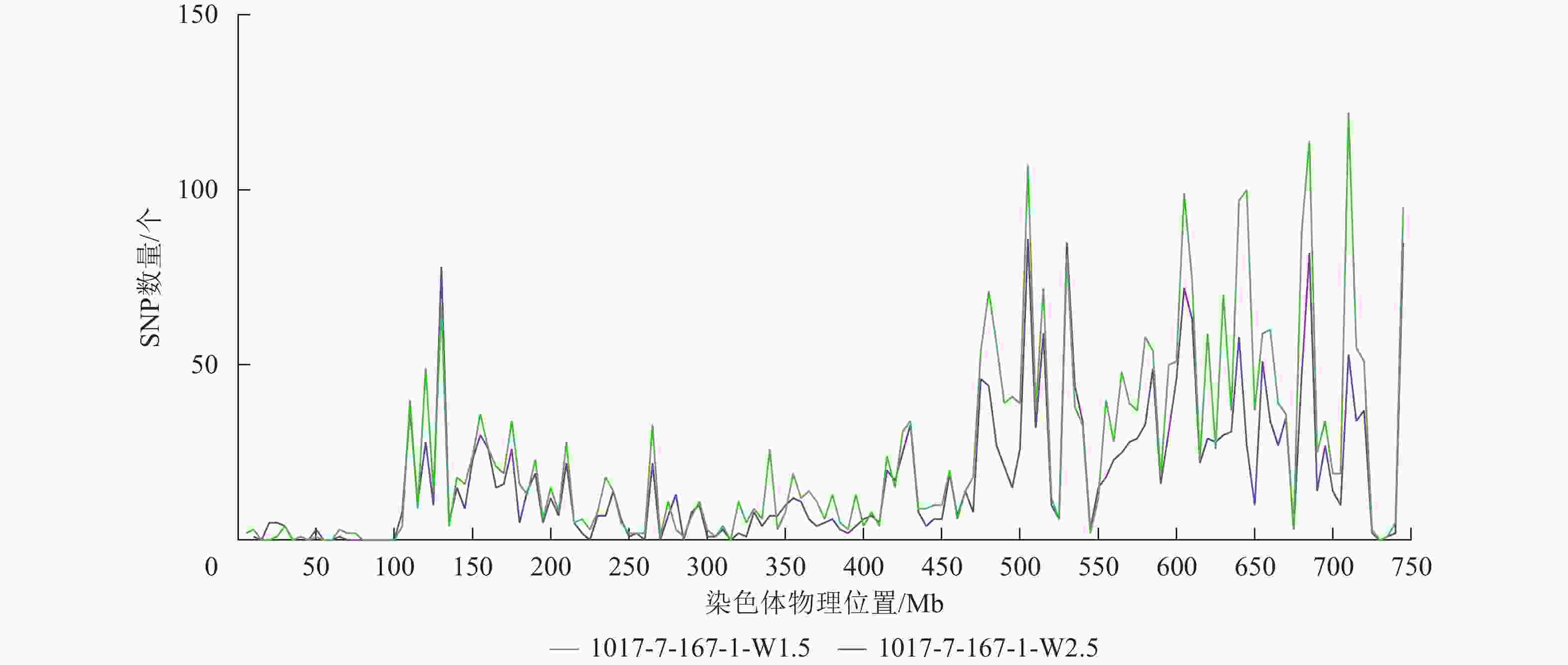
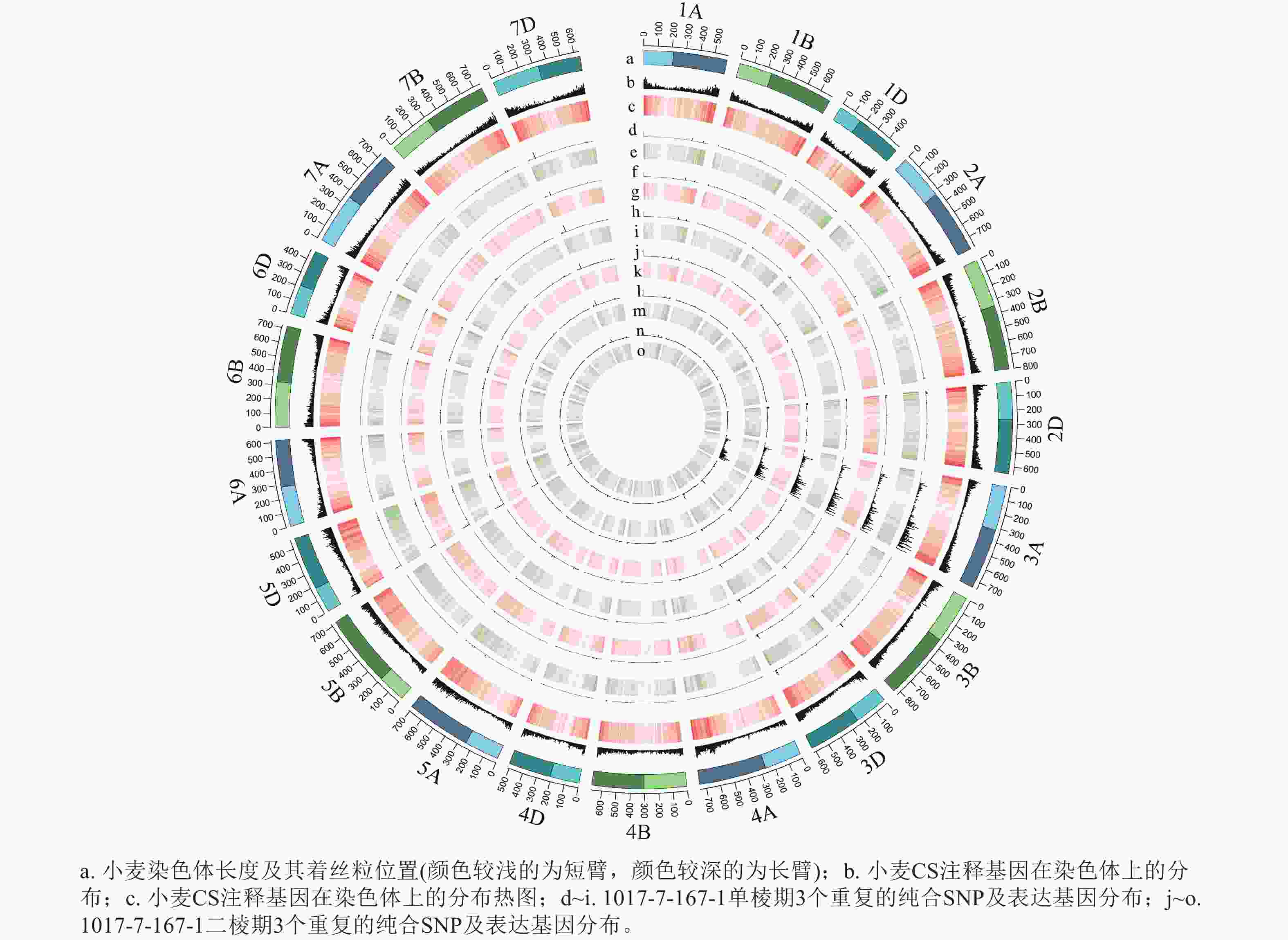
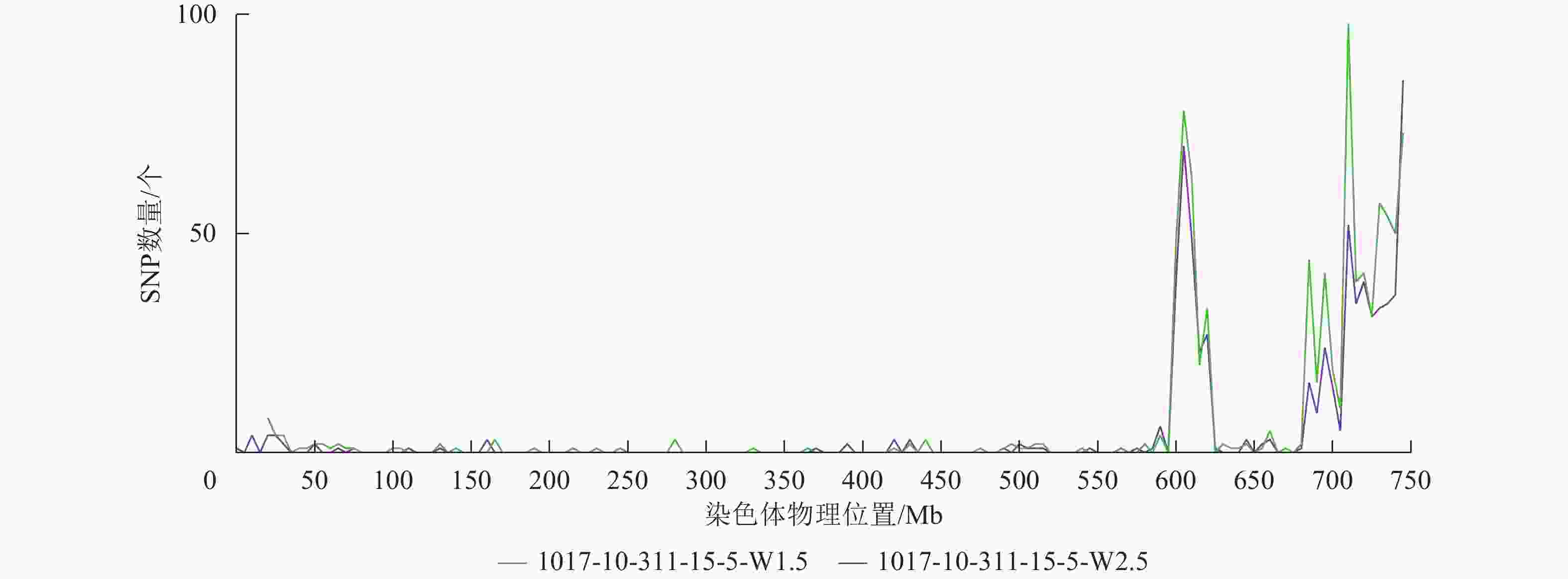
根据转录组测序数据,1017-7-167-1的纯合SNP主要分布在3A染色体的100~745 Mb处(图2~3),这表明该材料为CASL3AL的后代,并在3A染色体上携带了大片段的野生二粒小麦染色体片段。对于1017-10-311-15-5,其纯合SNP主要分布在3A染色体末端,位于600~745 Mb处(图4~5)。从染色体组成结果来看,这2个材料的遗传背景较为干净,所携带的野生二粒小麦染色体片段均主要位于3A染色体上。
-
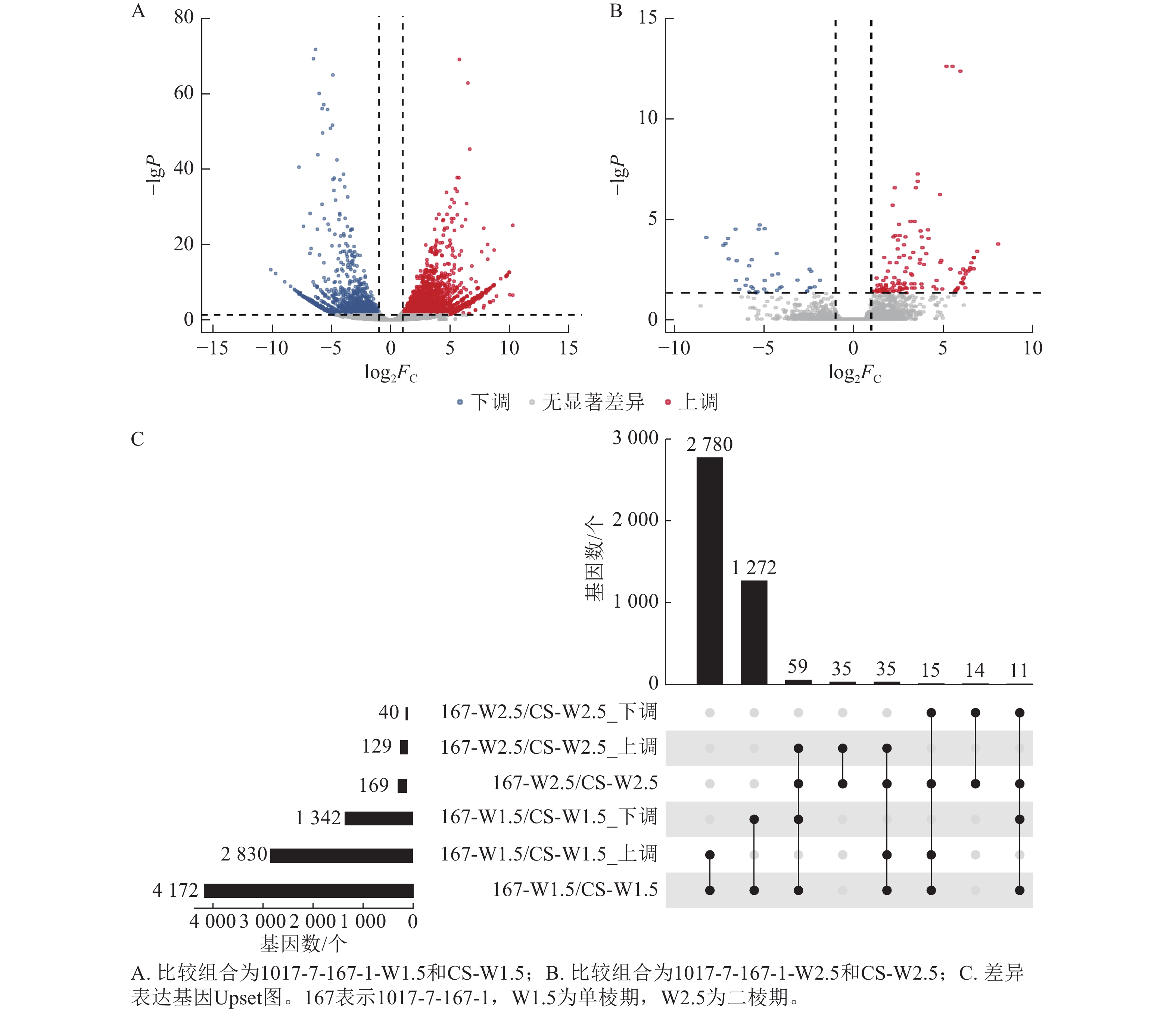
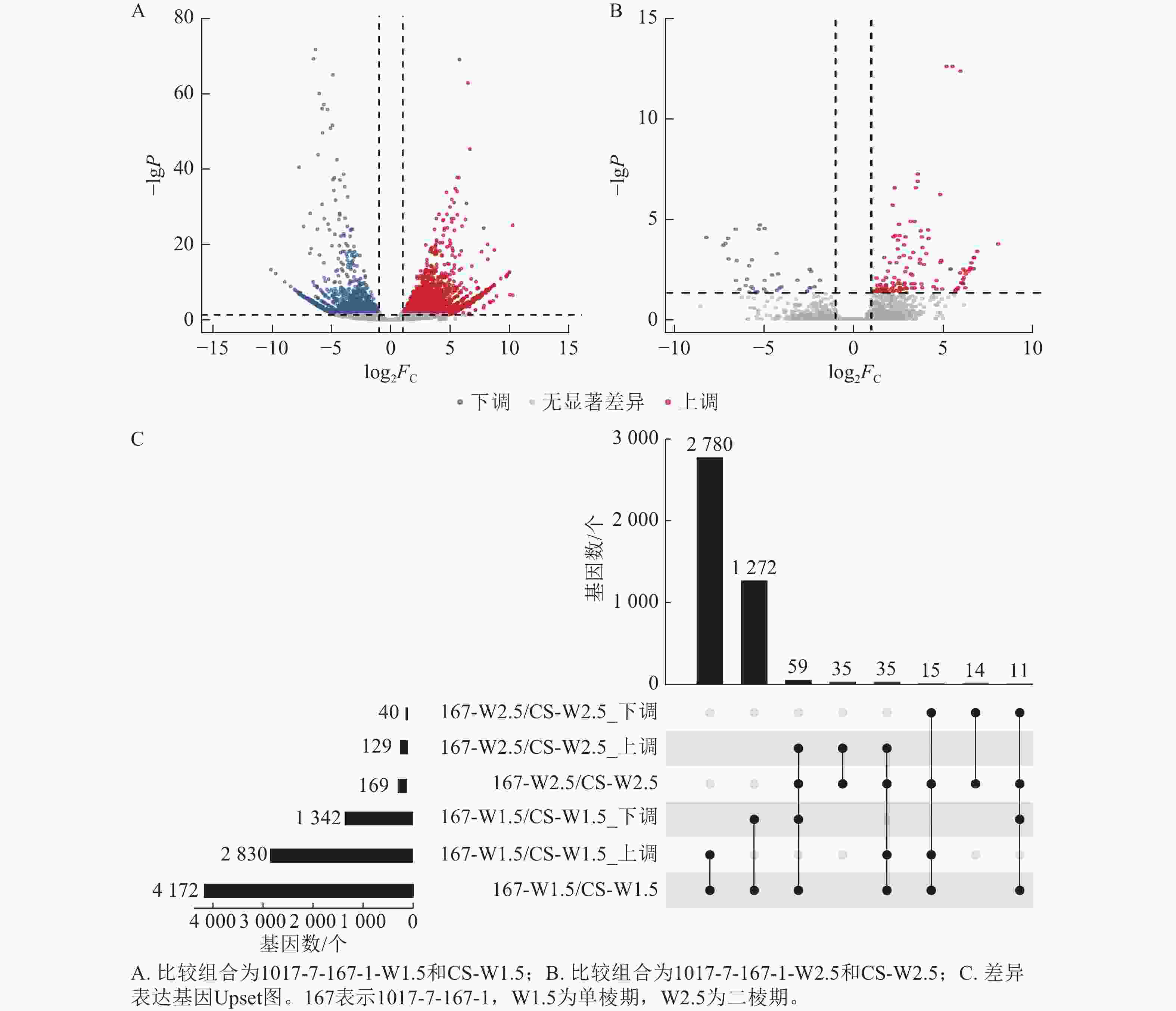
不同材料在相同时期的基因差异表达分析显示(图6):1017-7-167-1与CS相比,在单棱期的差异表达基因数量多于二棱期。具体而言,1017-7-167-1与CS相比,单棱期有2 830个基因上调表达,1 342个基因下调表达;而在二棱期,分别有129个基因上调表达和40个基因下调表达。其中,有35个基因在这2个时期都上调表达,11个基因在这2个时期都下调表达。总体来看,单棱期有4 172个基因差异表达,二棱期有169个基因差异表达。
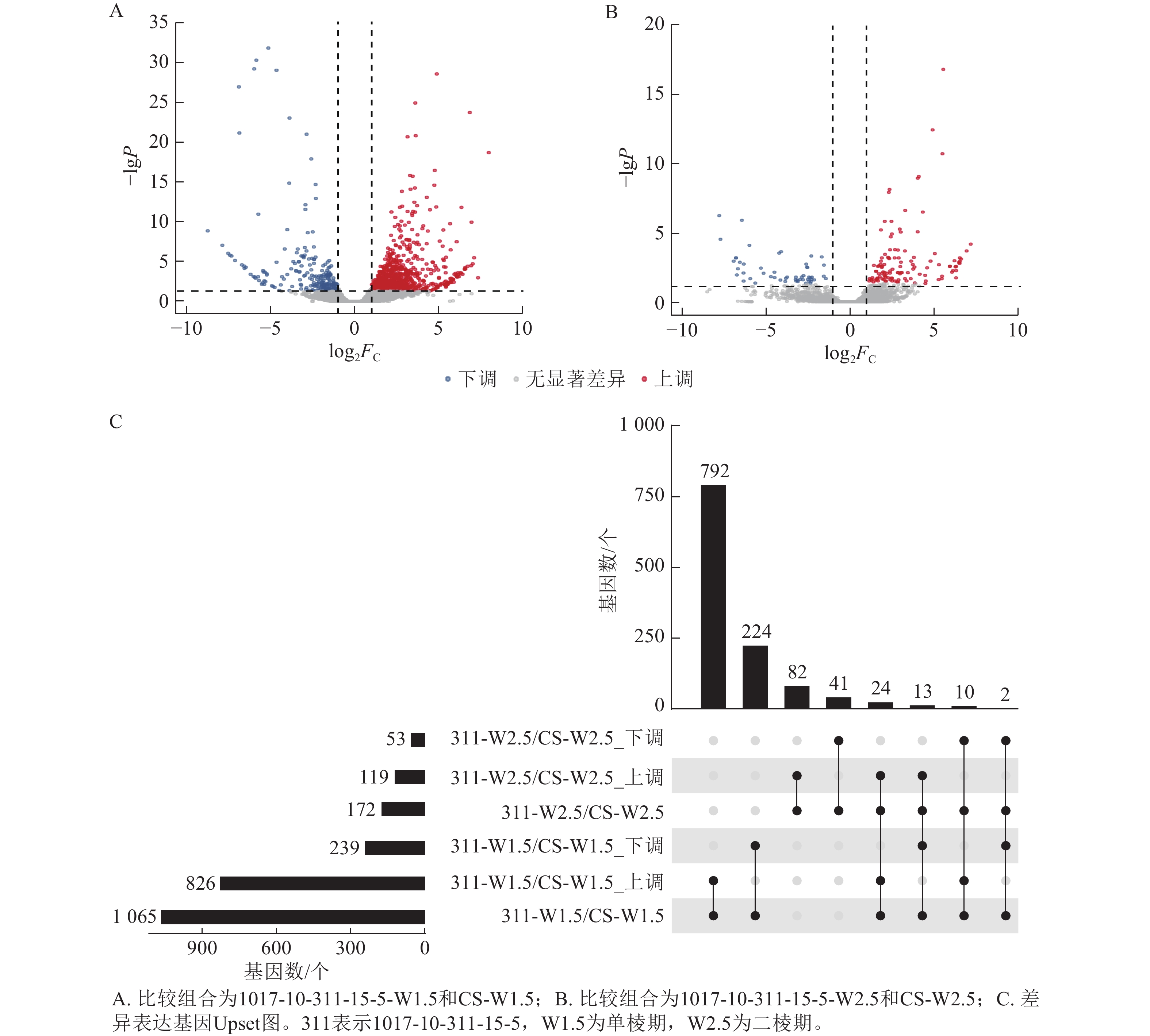
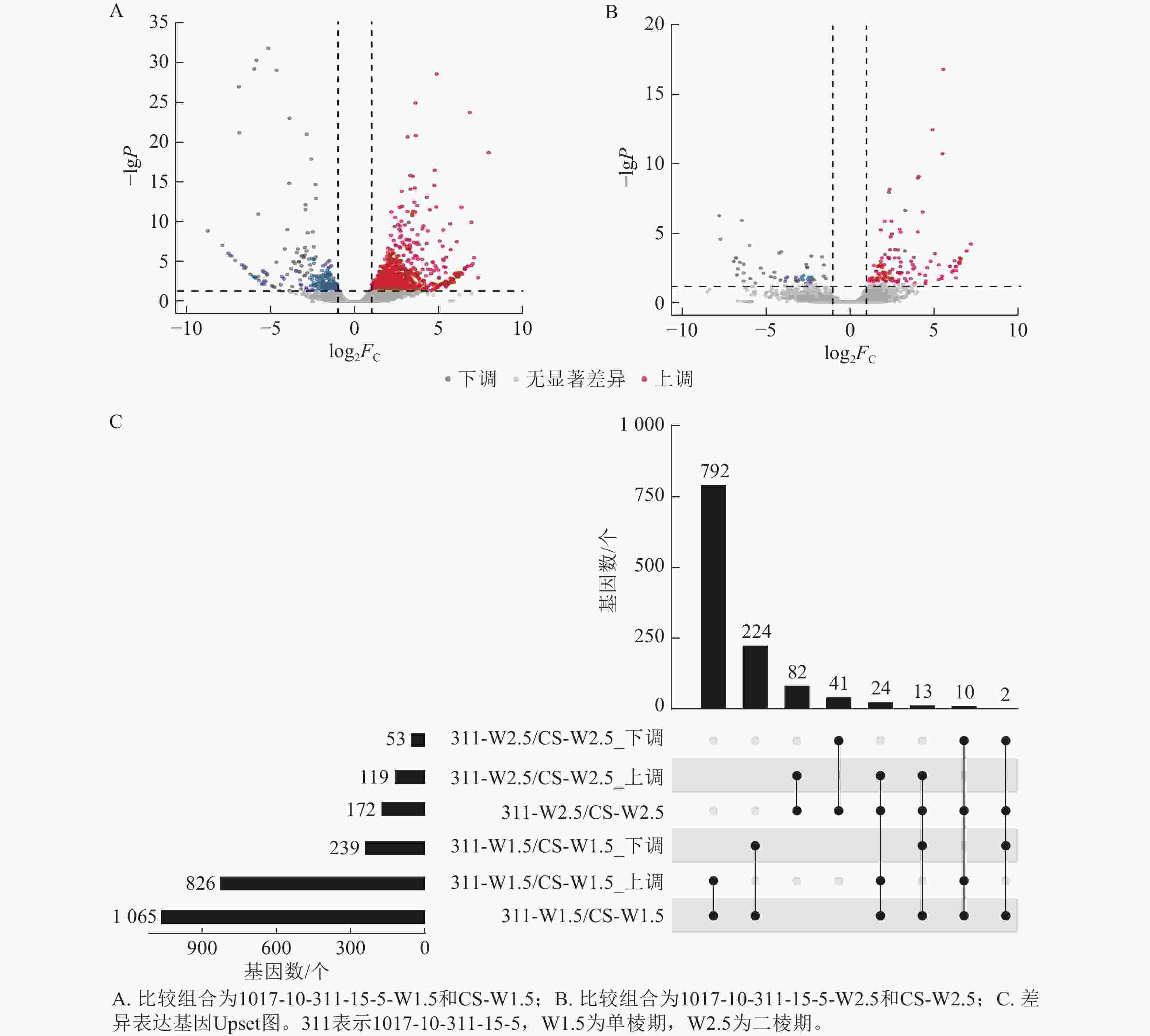
由图7可知:1017-10-311-15-5与CS相比,在单棱期有826个基因上调表达,239个基因下调表达;而在二棱期,分别有119个基因上调表达和53个基因下调表达。其中,有24个基因在这2个时期都上调表达,2个基因在这2个时期都下调表达。总体来看,单棱期有1 065个基因差异表达,二棱期有172个基因差异表达。
-
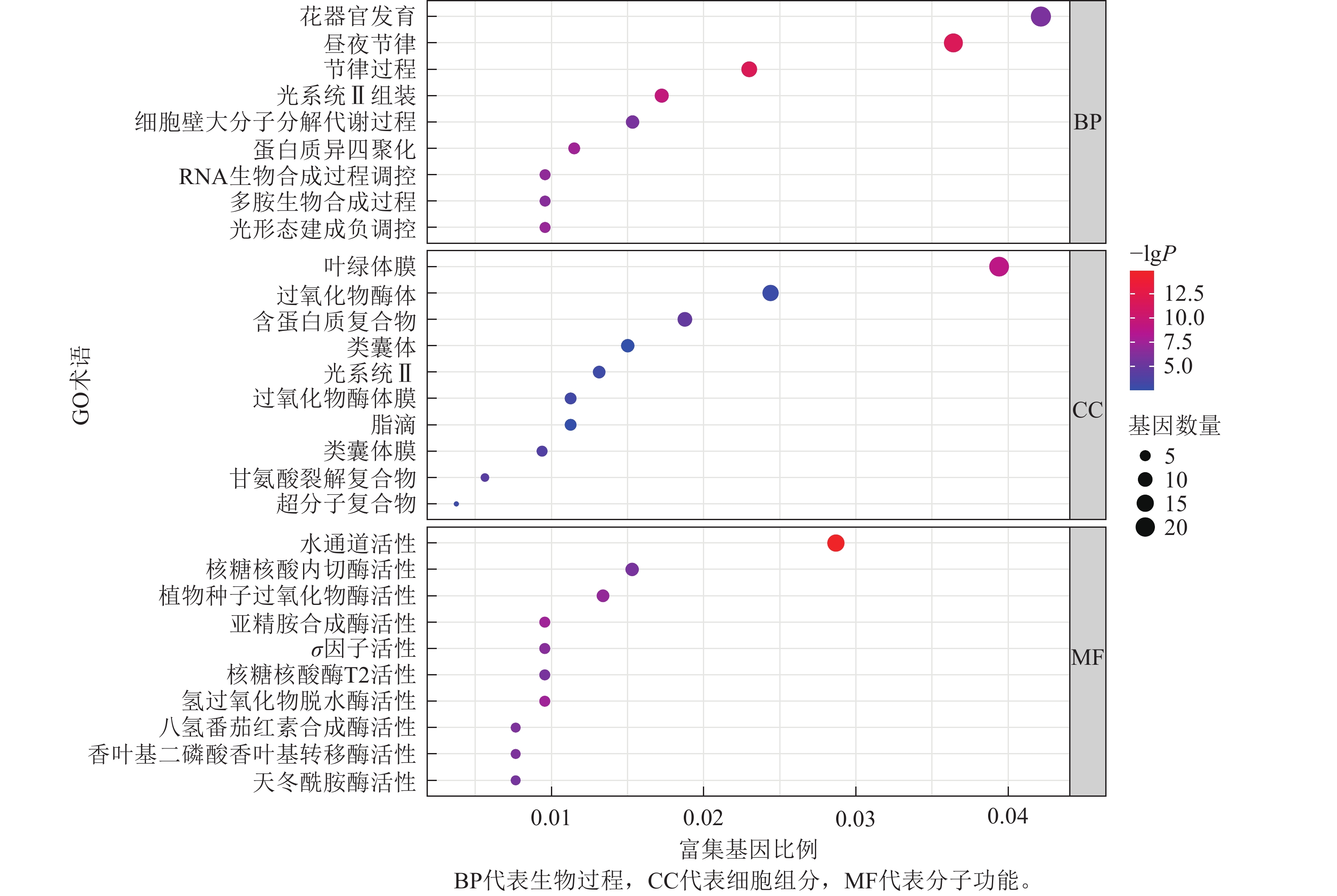
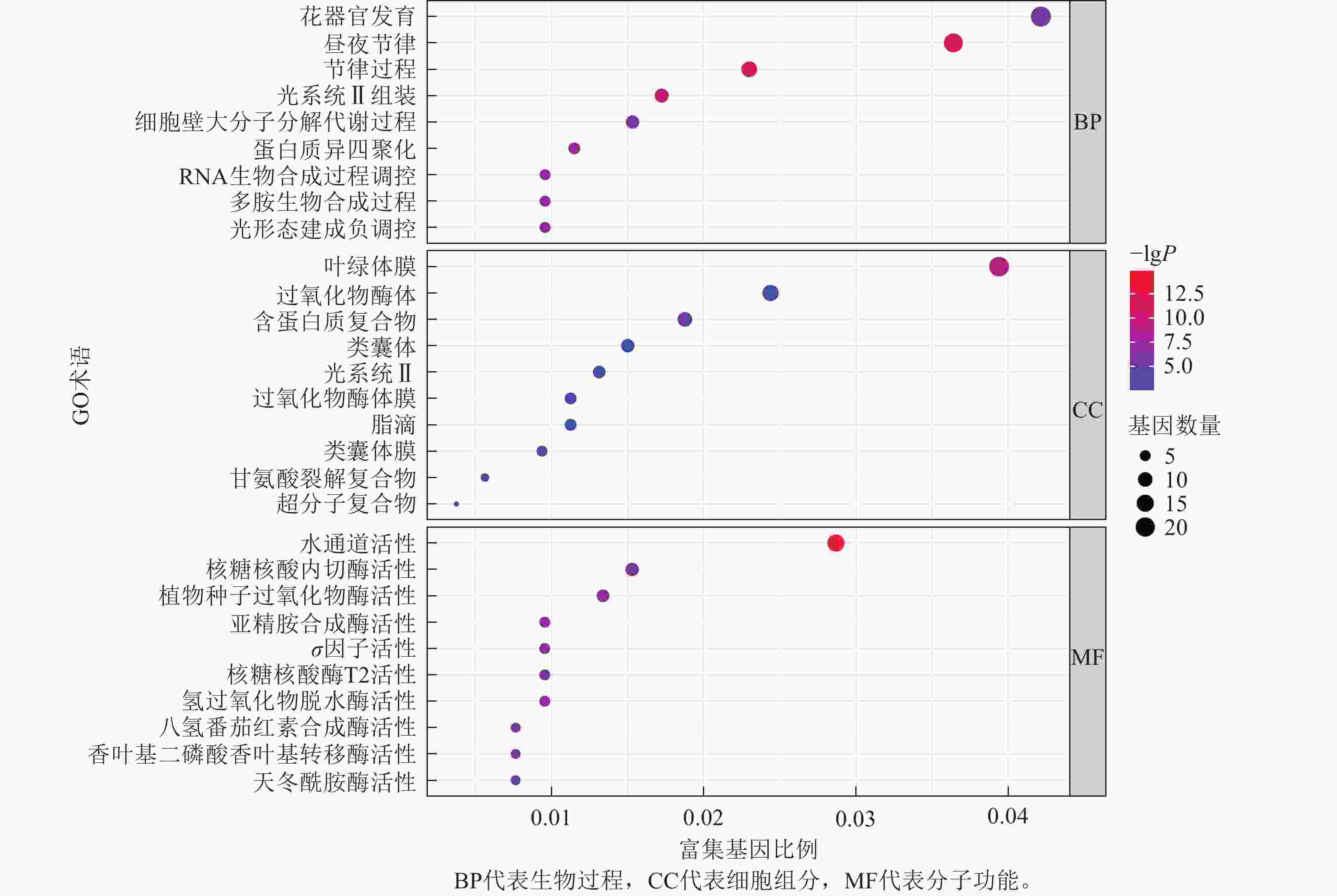
为了进一步探寻差异表达基因的功能,本研究对这些基因进行了GO富集分析,并将分析结果进行可视化展示(图8)。GO富集分析主要分为3类:生物过程(biological process,BP)、细胞组分(cellular component,CC)和分子功能(molecular function,MF)。
在生物过程方面,差异表达基因主要富集到3个最显著的GO术语为花器官发育,昼夜节律,节律过程。在细胞组分方面,差异表达基因主要富集到3个最显著的GO术语为叶绿体膜结构,过氧化物酶体,含蛋白质复合物。在分子功能方面,差异表达基因主要富集到3个最显著的GO术语为水通道活性,核糖核酸内切酶活性,植物种子过氧化物酶活性。
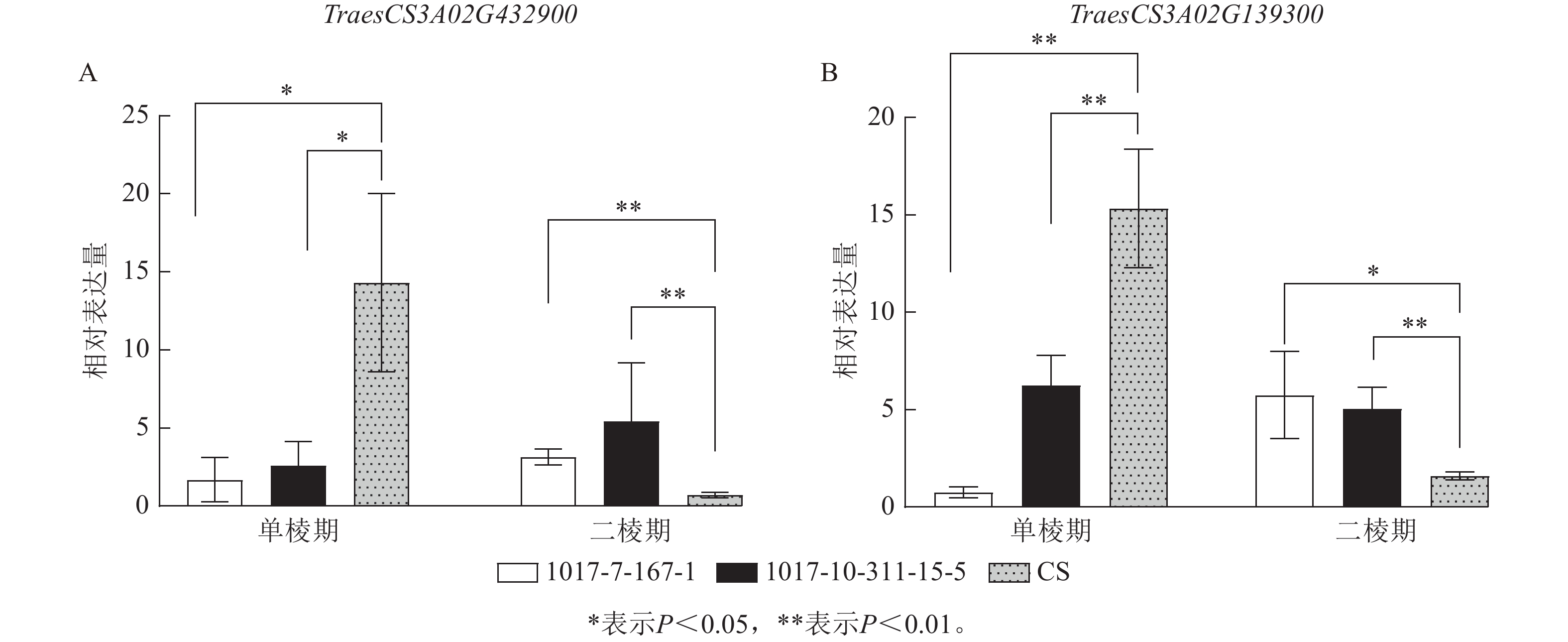
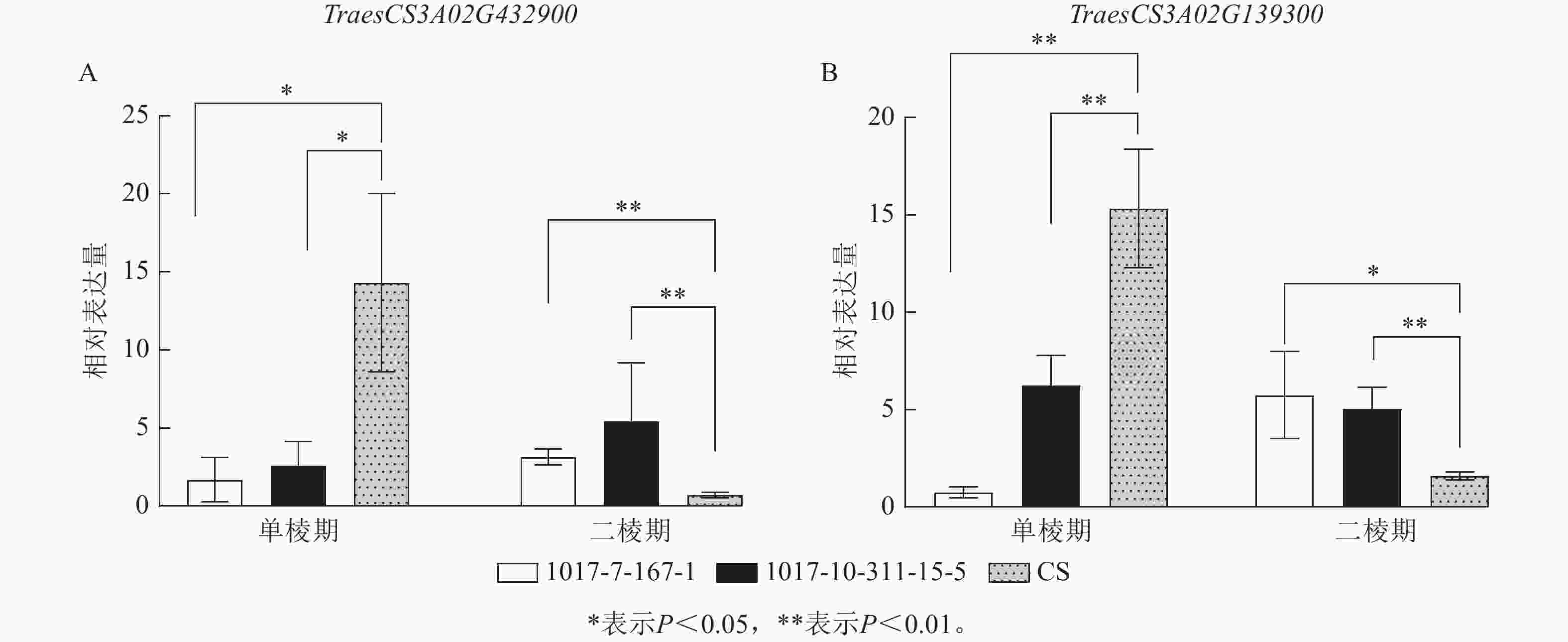
为进一步筛选调控CASL3AL早抽穗的候选基因,结合转录组测序的表达量结果,在3A染色体SNP富集区域筛选到了2个候选基因(图9)。其中TraesCS3A02G432900基因编码MADS-box转录因子,该转录因子主要参与调控植物开花时间以及花器官的形成。在单棱期,该基因在1017-7-167-1和1017-10-311-15-5中的表达量显著低于CS (P<0.05),而在二棱期,该基因在1017-7-167-1和1017-10-311-15-5表达量显著高于CS (P<0.01),因此推测该基因上调表达促进了1017-7-167-1和1017-10-311-15-5材料的幼穗发育进程,2个早熟材料相比于CS更早进入到二棱期,进而使得抽穗和开花时间也提前。TraesCS3A02G139300基因编码B-box锌指蛋白家族,该家族成员通过光周期、昼夜节律等途径影响植物开花时间,在单棱期,该基因在1017-7-167-1和1017-10-311-15-5的表达量显著低于CS (P<0.01),在生长至二棱期后,该基因在1017-7-167-1的表达量显著高于CS (P<0.05),在1017-10-311-15-5中的表达量也显著高于CS (P<0.01),因此推测该基因对于小麦幼穗发育具有正向调控作用。
-
小麦抽穗期作为开花之前的重要时期,是一个复杂的数量性状,受到环境和多种基因的共同调控[25]。野生二粒小麦作为普通小麦的祖先,拥有抗旱、抗病、耐盐、品质高等诸多优良性状,是现代小麦遗传改良育种领域的重要种质库和基因库[26−27]。CASL3AL是一种以CS为背景,3A染色体长臂被野生二粒小麦所置换的染色体臂置换系材料,在多元种植环境下均表现为早熟。本研究利用CASL3AL×CS后代中稳定早熟的材料1017-7-167-1和1017-10-311-15-5,以及晚熟的CS进行了RNA-Seq转录组分析,旨在更全面地挖掘野生二粒小麦染色体上促进普通小麦早抽穗的候选基因。
-
CASL3AL与CS进行杂交后,其后代经过多年种植和分离,1017-7-167-1材料的纯合SNP主要分布在3A染色体100~745 Mb处,而1017-10-311-15-5材料所包含的纯合SNP较少,集中在3A染色体末端600~745 Mb处。由于CASL3AL是将CS端体与野生二粒小麦杂交后,继续与CS端体进行多次的回交所产生的,因此CASL3AL×CS的后代材料在非置换臂上也会有少量的野生二粒小麦片段残留。这种遗传背景的复杂性为研究小麦抽穗期的遗传调控机制提供了丰富的材料基础。
-
本研究通过转录组测序和数据分析,在早熟材料与晚熟材料之间筛选出了差异表达基因。这些基因主要富集于GO条目的生物过程、细胞组分和分子功能,生物过程中包含花器官发育和昼夜节律等GO条目,与小麦的抽穗期密切相关,表明在抽穗期调控中,花器官发育和昼夜节律起着关键作用。
-
根据GO富集分析和转录组表达量的结果,筛选出了2个可能参与小麦抽穗期调控的关键基因。TraesCS3A02G432900编码MADS-box转录因子,该转录因子广泛参与植物的生长发育过程,在植物开花时间调控、花序结构、花器官形成等方面发挥着重要的作用[28]。小麦春化的关键基因Vrn1也属于MADS-box转录因子,在叶片和顶端分生组织中表达,随后促进开花[29]。TraesCS3A02G139300基因编码B-box锌指蛋白家族,是植物中一类重要的转录调控因子,主要通过参与光信号传导来影响开花时间[30]。在拟南芥Arabidopsis thaliana中,短日照和长日照条件下过表达含有B-box锌指结构域的STO基因,使得开花时间均提前[31]。在长日照低温条件下,拟南芥中的BBX28和BBX29锌指蛋白会与CO-FT调控模块协同作用以促进开花[32]。
-
本研究通过RNA-Seq转录组测序,系统分析了2个早熟材料的纯合SNP分布情况,筛选出了不同发育阶段的差异表达基因。这些基因富集于生物过程、细胞组分、分子功能、花器官发育、昼夜节律等GO条目中。同时筛选出了2个可能参与小麦开花调控的候选基因,这2个基因与MADS-box转录因子和B-box锌指结构域有关。后续可深入验证关键基因功能,探讨其环境响应表达调控机制,并继续挖掘幼穗发育时期富集到的差异表达基因,以期为小麦的遗传改良和育种提供更丰富的基因资源。
Candidate genes for early heading date in chromosome arm substitution line CASL3AL
doi: 10.11833/j.issn.2095-0756.20250260
- Received Date: 2025-04-19
- Accepted Date: 2026-01-09
- Rev Recd Date: 2025-12-01
- Available Online: 2026-04-02
- Publish Date: 2026-04-02
-
Key words:
- Triticum turgidum var. dicoccoides /
- chromosome arm substitution line /
- heading date /
- RNA-Seq /
- single nucleotide polymorphism (SNP) /
- wheat growth period improvement
Abstract:
| Citation: | DONG Jianhao, CHEN Liangpeng, ZHAO Yanhao, et al. Candidate genes for early heading date in chromosome arm substitution line CASL3AL[J]. Journal of Zhejiang A&F University, 2026, 43(2): 310−319 doi: 10.11833/j.issn.2095-0756.20250260 |









 DownLoad:
DownLoad: