-
AP2(APETALA2)/ERF(ethylene responsive factor,ERF)转录因子超家族是植物中重要的转录因子类型,其共同的结构特征是在DNA结合区包含1个或多个由约60~70个氨基酸组成的高度保守的AP2/ERF结构域。根据结构域数目及其所结合的顺式作用元件的差异,AP2/ERF转录因子超家族可分为:AP2,乙烯反应因子,干旱反应元件结合蛋白(dehydration responsive element binding protein,DREB)和RAV 4个转录因子家族。ERFs是植物中特有的转录因子家族,研究证实ERF转录因子参与调控包括植物生长、发育在内的多个生物学过程[1-3]。另外,通过AP2/ERF结构域和PR基因启动子顺式作用元件GCC-box(AGCCGCC)的结合,ERF基因参与植物对环境刺激,尤其是对病原菌胁迫的响应。为了发现AP2/ERF转录因子在林木抗非生物胁迫以及抗病性中的作用,近年来,一些研究者对多年生木本模式植物——杨树Populus spp.的ERF和DREB转录因子进行了深入研究。Nanjo等[4]在脱水、高盐、高温、冷冻、脱落酸以及过氧化氢处理后的意大利杨Populus nigra var. italica EST文库鉴定到13个ERF/AP2转录因子,其中一些仅在特定的环境胁迫下特异表达。Zhuang等[5]对杨树AP2/ERF转录因子超家族进行了全基因组分析,鉴别出了91个ERF转录因子以及77个DREB转录因子。Maki等[6]发现欧洲黑杨Populus nigra ERF基因——PncERF1基因在室温生长的杨树细胞内不表达,仅对低温处理后产生响应表达。Li等[7]报道转番茄Solanum lycopersicum ERF转录因子的杂交杨Populus alba × Populus berolinensis获得了对盐胁迫的抗性,并且在200.0 mmol·L-1氯化钠下,转基因杨的干物质总量、树高,叶片中脯氨酸含量、钠离子(Na+)含量均高于非转基因杨。最近,应用数字基因表达方法(digital gene expression,DGE),Chen等[8]发现了2个盐胁迫后下调表达的小黑杨Populus simonii × Populus nigra AP2/ERF转录因子。此外,研究发现ERF转录因子也在杨树顶芽的休眠诱导和维持过程中表达[9]。近来,作者在溃疡病菌胁迫的多种杨树杂交无性系中发现1个特定ERF转录因子基因(ERF-18基因)的上调表达,预期该基因可能与杨树的抗逆反应相关。为深入揭示该基因的功能,本研究对中国优良杨树乡土树种——山海关杨Populus deltoides ‘Shanghaiguan’ ERF-18基因(PdERF-18基因)进行生物信息学分析,实时定量PCR(real-time QPCR)技术检测其在不同环境胁迫下的表达模式,以期阐明该基因的功能和抗逆反应机制,并为杨树抗逆遗传育种提供实验基础。
-
植物材料:1年生山海关杨P23植株(中国林业科学研究院林业研究所)。杨树溃疡病菌:葡萄座腔菌Botryosphaeria dothidea菌株CZ060(保存于中国林业科学研究院林业新技术研究所),马铃薯葡萄糖琼脂(PDA)培养基(含马铃薯浸粉20.0 g·L-1,葡萄糖20.0 g·L-1,琼脂15.0 g·L-1),25 ℃下避光培养7 d。细菌病原:根癌农杆菌Agrobacterium tumefaciens(Rhizobium radiobacter)菌株GV3103,Luria-Bertani(LB)培养基(含胰蛋白胨10.0 g·L-1,酵母提取物5.0 g·L-1,氯化钠10.0 g·L-1),37 ℃下避光培养48 h。
-
从山海关杨枝干距地面20.0 cm起,以灭菌打孔器每隔15.0 cm在树皮上打孔10个·枝条-1,深度至露出木质部,直径为4.0 mm。将培养7 d的溃疡病菌接种至接种孔,接种部位以保鲜膜密封保湿,室温培养1,2,3,6,12,48,72 h后,刮取树皮(去除接种部位的树皮,芽点),液氮保存备用。以无菌PDA培养基接种杨树作为对照。利用十六烷基三甲基溴化铵(CTAB)法[10]提取溃疡病菌侵染的山海关杨总核糖核酸(RNA),Promega公司DNase消化,ND-8000核酸定量仪(NanoDrop公司,美国)检测RNA样品纯度,琼脂糖凝胶电泳检验RNA样品完整性。取2.0 μg总RNA样品,以PrimeScript Ⅱ第1链cDNA合成试剂盒(Takara公司,中国大连)进行反转录反应。以引物(forward引物5′-TTTATGTCGGCTGAGTTTGA-3′)和(reverse引物5′-GAG GGGTTTCTGTTGATGA-3′)在山海关杨P23植株中进行反转录-聚合酶链式反应(RT-PCR)扩增。PCR扩增体系包含1.0 μL cDNA,200.0 μmol·L-1 dNTPs,0.2 μmol·L-1 PCR引物,16.67 nkat Taq酶;扩增程序:94 ℃预变性1 min;94 ℃ 30 s,62 ℃ 30 s,72 ℃ 1 min,共35个循环;72 ℃延伸10 min。扩增产物以Promega公司T-easy载体克隆到大肠埃希菌Escherichia coli TOP10菌株,选择阳性克隆进行序列测定。
-
测序结果与毛果杨ERF基因进行比对分析,获得山海关杨ERF-18(PdERF-18)基因的编码序列(CDS),通过MEGA v5.0分析软件[11]翻译为氨基酸序列,应用在线分析平台ExPASy http://web.expasy.org/compute_pi/计算蛋白质分子量和预测编码蛋白等电点。
-
依据ERF转录因子ERF/AP2结构域的氨基酸序列,参照Sakuma等[12]的AP2/ERF转录因子的分类方法,应用MEGA软件[11]对PdERF-18基因进行系统发育分析。参比序列为12个拟南芥Arabidopsis thaliana ERF转录因子序列。这些序列分属6个ERF类群。多序列比对采用MUSCLE软件[13],系统发育分析方法为邻位连接法(neighbor joining,NJ),Bootstrap 1 000次检验进化树各分支的支持率。
-
①激素处理。分别以20.0 μmol·L-1水杨酸溶液(salicylic acid,SA),10.0 μmol·L-1茉莉酸溶液(jasmonic acid,JA)喷雾处理1年生山海关杨叶片,1,3,6,12 h后取叶片,液氮保存后备用。无菌水处理的植株作为对照。②非生物胁迫处理。1)脱水处理:山海关杨植株小心从培养容器中取出,水洗去掉泥土,吸干根部水分后置于定性滤纸上,25 ℃下放置1,2,3,6,12 h后取叶片。以无菌水喷洒并作保湿处理的植株作为对照。2)高温处理:将山海关杨在42 ℃培养箱中培养1,2,3,6,12 h后取叶片备用。室温培养的材料作为对照。3)盐胁迫:以100.0 mL浓度为2.3 mol·L-1氯化钠的水溶液浇注山海关杨根部,处理1,3,6,12 h后取叶片备用。4)叶片机械损伤:在山海关杨叶片上以直径4.0 mm的打孔器打孔3个·处理-1,处理1,2,3,6,12,24,48 h后取叶片备用。③生物胁迫。1)杨树溃疡病菌侵染:接种方法同1.2.1。2)根癌农杆菌侵染:从山海关杨枝干距地面20.0 cm起,隔15.0 cm刺伤杨树树皮(在5.0 mm的范围内刺伤5个点),共刺伤10个部位。以灭菌木签蘸取培养72 h的根癌农杆菌菌落,接种于针刺点,接种完成后以保鲜膜封闭保湿,室温下培养1,2,3,6,12,48,72 h后,刮取树皮(去除接种部位的树皮),液氮保存备用。以刺伤未接种的材料作为对照。以上试验中,实验材料、对照均为3株·处理-1。对以上处理进行总RNA提取,RNA提取方法同2.1.2。④生物及非生物胁迫下,PdERF-18基因的表达分析,采用Takara公司TAKARA SYBR Premix Ex Taq(DRR041)(Perfect Real Time)试剂盒(Takara公司,中国大连)进行山海关杨ERF-18基因表达的荧光定量PCR检测。RT-qPCR在ABI Prism 7500 HT序列检测系统上(AB公司,美国)上完成,两步法扩增。RT-qPCR检测引物为5′-CTTCTGTTC TTGAACTTGAC-3′和5′-CAAAGGATCGGTCAT AA AAG-3′。反应过程如下:50 ℃ 2 min,95 ℃ 10 min,95 ℃ 15 s,60 ℃ 1 min,共40个循环;通过扩增曲线以及琼脂糖凝胶电泳检测扩增反应的特异性。采用生物学重复2次·样品-1,采用技术重复6个·反应-1。相对基因表达量计算采用2-⊿⊿CT法[14],以UBQ基因(forward引物5′-GTTGATTTTTGCTGGGAAGC-3′;reverse引物5′-GATCT TGGCCTTCACGTTGT-3′)作为内参基因。
-
通过RT-PCR方法,在溃疡病菌侵染的山海关杨中克隆到1个ERF转录因子基因(PdERF-18),测序结果显示:该序列包含完整的CDS序列[美国生物技术信息中心(NCBI)序列登录号:JX275831],大小为936 bp,编码312个氨基酸残基。对比CDS序列和DNA测序结果,显示PdERF-18基因不编码内含子。
结构域分析显示,PdERF-18基因编码蛋白N端第89~137个氨基酸残基区域为ERF转录因子所特有的AP2/ERF保守结构域。AP2/ERF结构域中,与ERF转录因子分类及功能密切相关的第14编码、第19编码位点分别为丙氨酸(Ala,A)和谷氨酸残基(Glu,E)。另外,N端72~88位点包含有由碱性氨基酸残基组成的ERF转录因子核定位信号KRRR。而在C端则存在由大量酸性氨基酸残基组成的保守序列,这可能是ERF蛋白的转录激活域。该区域可能与胁迫条件下,下游基因表达的激活有关[12]。
ExPASy软件分析表明PdERF-18基因编码蛋白质的分子量(MW)为34 463.2 u,蛋白质表面具有较低电势,其等电点(PI)为4.63。
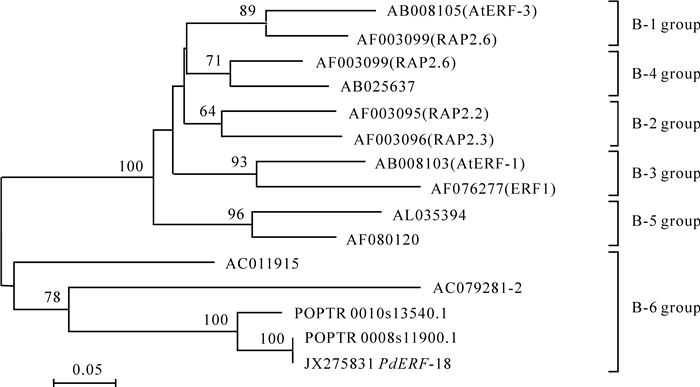
在杨树基因组数据库进行的Blast分析结果显示:PdERF-18基因与毛果杨ERF转录因子基因(POPTR_0008s11900.1和POPTR_0010s13540.1)高度同源。PdERF-18和POPTR_0008s11900.1的遗传距离最小,仅为0.013,在312个氨基酸位点中,仅有6个氨基酸残基发生了替换。POPTR_0008s11900.1基因同样不包含内含子。因此,我们推测PdERF-18是POPTR_0008s11900.1的直系同源基因。
以AP2/ERF结构域完成的系统发育分析显示,山海关杨PdERF-18,POPTR_0008s11900.1和POPTR_0010s13540.1均属于ERF转录因子B-6类群(图 1)。
-
实验结果表明:水杨酸和茉莉酸处理均能诱导山海关杨叶部PdERF-18基因表达上调(图 2)。水杨酸处理后1~3 h,PdERF-18基因表达水平不变;处理后6 h,PdERF-18基因表达水平为对照的2.60倍;而在处理后12 h,PdERF-18基因表达水平为对照的4.92倍(图 2A)。在茉莉酸处理中,PdERF-18基因表达在3 h时达到峰值,其相对表达量是对照的5.48倍,随后基因表达水平逐渐下降;处理后12 h,PdERF-18基因的表达量仅为对照的12%(图 2B)。结果显示:PdERF-18基因对于水杨酸和茉莉酸的反应模式并不完全相同。在处理后1~12 h,茉莉酸诱导PdERF-18基因迅速到达表达高峰,随后表达下降,甚至出现抑制表达;而水杨酸诱导的基因表达水平则在不断增加。
-
脱水、高温、盐胁迫甚至机械创伤均可诱导山海关杨叶部PdERF-18基因的特异性表达。如图 3所示,在这4种不同的环境胁迫下,处理初期(1~2 h)PdERF-18基因基因表达水平不变或略有下降;处理后3~6 h,各处理样品中PdERF-18基因的表达均大幅度上升。如,脱水、盐胁迫和机械创伤6 h后,PdERF-18基因表达均达到最高峰,其相对表达量分别为对照的15.90,9.18和8.72倍;而高温处理3 h时PdERF-18基因表达达到最高峰,其表达水平为对照的8.91倍,高温处理6 h时,PdERF-18基因表达水平为对照的3.04倍。非生物胁迫处理6 h后,脱水和高温处理样品中PdERF-18基因几乎不表达(相对表达量小于对照样品10%),盐胁迫和机械创伤样品中基因表达则和对照表达水平相近;对于机械创伤样品,在处理后48 h,仍能检测到一定的基因表达。
-
研究结果表明:葡萄座腔菌和根癌农杆菌处理均能诱导山海关杨PdERF-18基因的上调表达。在处理后1~48 h,葡萄座腔菌侵染样品的PdERF-18基因表达水平呈现近似正态曲线分布,处理后6 h,基因表达达到顶峰,其表达水平为对照的5.12倍(图 4A)。在本研究1~48 h的7个检测点,根癌农杆菌侵染样品中的PdERF-18基因表达水平均大于对照样品;在3~48 h的时间范围内,PdERF-18基因表达水平增加1.50倍以上,最高达到对照的3.14倍(图 4B)。
-
AP2/ERF结构域,尤其是AP2/ERF结构域的第14和第19位氨基酸残基对AP2/ERF转录因子的分类以及功能具有决定性的影响。DREB转录因子的典型特征是AP2/ERF结构域中第14位缬氨酸(V14)和第19位谷氨酸(E19)残基的存在。这2个位点的氨基酸残基替换(缬氨酸替换为丙氨酸,或者谷氨酸替换为天冬氨酸)均显著降低了突变型的DREB分子与DRE作用元件(TACCGACAT)的结合能力[12],并最终影响下游基因的诱导表达。而植物ERF转录因子的典型特征则是第14位丙氨酸(A14)和第19位天冬氨酸(D19)的存在[12]。例如在全部176个毛果杨Populus trichocaroa ERF转录因子(植物转录因子数据库PlantTFDB v2.0,)中,有173个ERF具有以上特征。然而,本研究发现,PdERF-18基因、杨树POPTR_0008s11900.1和POPTR_0010s13540.1基因编码产物的AP2/ERF结构域的第14位丙氨酸保守存在,第19位天冬氨酸则被酸性更强的谷氨酸(E19)所替换。显而易见,这种替换会导致蛋白质分子表面电势的变化,并可能会影响ERF分子与其顺式作用因子的结合能力。因此,我们推测PdERF-18转录因子可能具有与其他典型ERF转录因子不同的特点;或者由于第19位氨基酸残基的替换,PdERF-18转录因子具有结合其他顺式作用因子的能力。
-
植物主要通过诱导型基因的表达而对各种环境胁迫发生响应,而等植物诱导型基因(如PR蛋白基因、NBS-LRR蛋白基因等)的表达则受到特定转录因子在转录水平上的调控[15]。鉴于ERF转录因子在植物抗病、抗逆反应以及生理代谢中的重要作用,ERF转录因子基因的分离与功能研究成为了植物学研究的热点之一。McGrath等[16]提出了有效分离克隆ERF基因2步法策略:即首先鉴定出在病原诱导早期表达增强的转录因子基因,然后对候选的转录因子基因进行转基因、功能鉴定。考虑到胁迫处理后,ERF基因可能具有的动态表达特征,处理样品的采集时间对于转录因子的初步筛选具有非常重要的影响。
研究表明:尖镰孢菌Fusarium oxysporum接种6 h的样品适宜于拟南芥ERF基因的初筛[16],而高温、低温、脱落酸、萘乙酸、吲哚乙酸和赤霉素处理后,毛白杨DREB转录因子基因的最高表达出现在处理后6 h,干旱和盐胁迫下最高表达出现在处理后5 h和24 h[17]。而本研究结果显示:除根癌农杆菌外,其他7种生物或非生物胁迫均能诱导PdERF-18基因在处理后6 h高量表达,并且其表达水平均为对照的2.00倍以上。基于以上的结果,我们认为应用两步法分离克隆ERF基因时,胁迫处理后6 h是样品采集的适宜时间。
-
植物抗病反应过程中,病菌侵染信号的传递主要通过水杨酸(SA)信号传导途径和茉莉酸(JA)/乙烯(ET)途径完成。在拟南芥Arabidopsis thaliana中,2个信号转导途径参与对不同类型病原菌的反应。SA途径主要在活体营养型病原菌的“Gene-for-Gene”抗性和系统抗性中发挥相关作用[18]。JA/ET途径主要参与寄主多基因控制的对腐生型真菌、土传真菌、卵菌等的抗性反应[19-21]。杨树中,JA/ET途径与机械损伤反应、黑斑病菌Marssonina brunnea f. sp. multigermtubi侵染有关,在诱导杨树间接、直接防御能力的过程中起着关键作用[22-24]。
杨树溃疡病是由葡萄座腔菌引起的中国北方地区主要林木病害,其病原为典型的弱寄生性真菌,同时具有一定内生性。那么,杨树通过哪种信号转导途径对葡萄座腔菌侵染产生抗性反应呢?本研究中,病原菌接种以及激素处理后ERF基因的表达模式为我们了解杨树病害发生过程中的激素信号转导提供了新的视角。如图 2B和图 4A所示:溃疡病菌侵染和茉莉酸处理后,PdERF-18转录因子表达的变化趋势具有一致性,即在处理后基因表达水平逐渐上升,处理后3 h或6 h表达量最高,随后表达水平逐渐降低,甚至达到对照水平之下。据此,我们推测山海关杨对于溃疡病菌侵染和茉莉酸具有较为相近的应激反应,而杨树溃疡病发生中的信号转导可能与JA/ET途径途径有较大相关性。
根癌农杆菌不仅是重要植物遗传转化载体,也是常见的土传细菌性病原,可以引起包括杨树、桃树Prunus presica 等在内的许多植物的根茎部肿瘤。研究证明:植物根癌病的发生是根癌农杆菌携带的毒性基因与植物的抗性基因之间亲和性互作的结果[25-28]。本研究结果显示:水杨酸处理和根癌农杆菌侵染后山海关杨PdERF-18基因的表达也表现出较为一致的变化趋势。因此,水杨酸信号传导通路可能在杨树根癌病的信号传递过程中起着重要作用。然而,由于实验设计的原因,我们仅仅检测了水杨酸处理后0~12 h PdERF-18基因的表达,12 h之后水杨酸是否保持持续上调表达的趋势,还需要在今后的研究中进行深入的分析。
另外,很多ERF转录因子是水杨酸、茉莉酸和乙烯信号转导途径交叉处的重要成分,居于植物基因调控网络的节点位置[29-30]。本研究结果显示:PdERF-18基因的表达不仅与多种生物及非生物环境胁迫相关,也与水杨酸和茉莉酸激素信号相关,因此,我们推测PdERF-18也可能处于杨树基因调控网络的节点位置。通过该基因的表达,杨树对机械损伤、低温、脱水、盐胁迫以及溃疡病菌和根癌农杆菌侵染产生响应。对山海关杨PdERF-18基因表达及功能的研究,可以深化对杨树抗逆境胁迫能力的认识,并为杨树抗逆遗传转化提供新的基因资源。
Expression patterns for a PdERF-18 response to different stresses in Populus deltoides 'Shanhaiguan'
-
摘要: 对一个溃疡病菌Botryosphaeria dothidea胁迫相关的山海关杨Populus deltoides ‘Shanghaiguan’乙烯反应因子(ethylene responsive factors, ERFs)基因——PdERF-18基因的系统发育、编码产物结构以及多种胁迫下的表达模式进行了分析。结果表明:PdERF-18基因编码产物属于ERF转录因子家族B-6类群, 具有植物ERF转录因子的典型结构, 但它与转录调控功能密切相关的AP2/ERF结构域第19位氨基酸位点发生替换, 推测PdERF-18基因可能具有不同于其他典型ERF转录因子的功能; 实时定量分析表明:溃疡病菌、根癌农杆菌Agrobacterium tumefaciens(Rhizobium radiobacter), 脱水、机械胁迫、盐胁迫和高温处理、茉莉酸以及水杨酸处理均可诱导PdERF-18基因的上调表达。其中溃疡病菌与茉莉酸处理以及根癌农杆菌与水杨酸处理下, 杨树PdERF-18基因分别具有相似的表达特征, 显示溃疡病菌、根癌农杆菌侵染的信号转导可能分别与茉莉酸/乙烯途径、水杨酸信号传导途径有关。PdERF-18基因可以对多种非生物、生物胁迫产生响应, PdERF-18基因有可能作为一种新的遗传资源而应用于杨树抗性遗传改良。Abstract: Ethylene responsive factors (ERFs), one kind of APETALA2(AP2)/ERF transcriptional factor that responds to environmental stimuli, especially to pathogen attacks, play an important role in plant-pathogen interactions. Recently, we noticed that one ERF-18 gene involved in the pathogenesis processes of the poplar stem canker disease of some poplar hybrid clones. However, the expression patterns of this gene response to other environmental stimuli are unclear. In this study, the phylogenesis, structure, and expression patterns of one Populus deltoides ERF gene (PdERF-18) were reported. Treatments including salicylic acid and jasmonic acid (JA) induction, Botryosphaeria dothidea and Agrobacterium tumefaciens (Rhizobium radiobacter) inoculation, mechanical stress, high temperature, salt stress, and dehydration stress were tested with real-time reverse transcription polymerase chain reaction (RT-qPCR). Results revealed a PdERF-18 coding for a B-6 member in the ERF family B. The 14th alanine and 19th aspartate residues in the AP2/ERF domain, typically characteristic of plant ERFs, were altered with an aspartate site substituted by a glutamate (an amino acid residue with stronger electronegativity than aspartate) in the coding product for the PdERF-18 gene implying that PdERF-18 might have some additional functions besides its resistance to a pathogen. RT-qPCR data of the diverse stresses revealed up-regulated expressed patterns in P. deltoides. Also, similar expression pattern responses for both B. dothidea and JA as well as responses to A. tumefaciens and salicylic acid were found. Thus, it was inferred that the JA/ethylene (ET) transduction pathway regulated gene expression in poplar canker pathogen attacks, but the SA pathway was involved in the responsiveness to poplar crown gall disease; therefore, it was deduced that PdERF-18 was a multi-functional transcriptional factor involved in many biotic and abiotic stresses.
-
-
[1] FENG Jianxun, LIU Di, PAN Yi, et al. An annotation update via cDNA sequence analysis and comprehensive profiling of developmental, hormonal or environmental responsiveness of the Arabidopsis AP2/EREBP transcription factor gene family[J]. Plant Mol Biol, 2005, 59(6):853-868. [2] GUTTERSON N, REUBER T L. Regulation of disease resistance pathways by AP2/ERF transcription factors[J]. Curr Opin Plant Biol, 2004, 7(4):465-471. [3] NAKANO T, SUZUKI K, FUJIMURA T, et al. Genome-wide analysis of the ERF gene family in Arabidopsis and rice[J]. Plant Physiol, 2006, 140(2):411-432. [4] NANJO T, FUTAMURA N, NISHIGUCHI M, et al. Characterization of full-length enriched expressed sequence tags of stress-treated poplar leaves[J]. Plant Cell Physiol, 2004, 45(12):1738-1748. [5] ZHUNAG Jing, CAI Bin, PENG Rihe, et al. Genome-wide analysis of the AP2/ERF gene family in Populus trichocarpa[J]. Biochem Biophys Res Commun, 2008, 371(3):468-474. [6] MAKI H, SATO M, OGAWA K, et al. Cloning and expression profile of an ERF gene isolated from cold-stressed poplar cells (Populus nigra)[J]. Cytol Int J Cytol, 2011, 76(1):11-18. [7] LI Yiliang, SU Xiaohua, ZHANG Bingyu, et al. Expression of jasmonic ethylene responsive factor gene in transgenic poplar tree leads to increased salt tolerance[J]. Tree Physiol, 2009, 29(2):273-279. [8] CHEN Su, JIANG Jing, LI Huiyu, et al. The salt-responsive transcriptome of Populus simonii×Populus nigra via DGE[J]. Gene, 2012, 504(2):203-212. [9] ROHDE A, RUTTINK T, HOSTYN V, et al. Gene expression during the induction, maintenance, and release of dormancy in apical buds of poplar[J]. J Exper Bot, 2007, 58(15/16):4047-4060. [10] CHANG Shujun, PURYEAR J, CAIRNEY J. A simple and efficient method for isolating RNA from pine trees[J]. Plant Mol Biol Rep, 1993, 11(2):113-116. [11] TAMURA K, PETERSON D, PETERSON N, et al. MEGA5:molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods[J]. Mol Biol Evol, 2011, 28(10):2731-2739. [12] SAKUMA Y, LIU Qiang, DUBOUZET J G, et al. DNA-binding specificity of the ERF/AP2 domain of Arabidopsis DREBs, transcription factors involved in dehydration and cold inducible gene expression[J]. Biochem Biophys Res Commun, 2002, 290(3):998-1009. [13] EDGAR R C. MUSCLE:multiple sequence alignment with high accuracy and high throughput[J]. Nucl Acids Res, 2004, 32(5):1792-1797. [14] LIVAK K J, SCHMITTGEN T D. Analysis of relative gene expression data using real-time quantitative PCR and and the 2-⊿⊿CT method[J]. Methods, 2001, 25(4):402-408. [15] AGARWAL M, HAO Yujin, KAPOOR A, et al. A R2R3 type MYB transcription factor is involved in the cold regulation of CBF genes and in acquired freezing tolerance[J]. J Biol Chem, 2006, 281(49):37636-37645. [16] McGRATH K C, DOMBRECHT B, MANNERS J M, et al. Repressor and activator type ethylene response factors functioning in jasmonate signaling and disease resistance identified via a genome-wide screen of Arabidopsis transcription factor gene expression[J]. Plant Physiol, 2005, 139(2):949-959. [17] 郭琦, 王保垒, 王博文, 等.毛白杨PtDREB2A基因的克隆、表达及单核苷酸多态性分析[J].林业科学, 2011, 47(4):49-56. GUO Qi, WANG Baolei, WANG Bowen, et al. Isolation, expression and single nucleotide polymorphisms analysis of PtDREB2A in Populus tomentosa[J]. Sci Silv Sin, 2011, 47(4):49-56. [18] 刘士旺, 吴学龙, 郭泽建.拟南芥的抗病信号传导途径[J].植物病理学报, 2003, 33(2):104-111. LIU Shiwang, WU Xuelong, GUO Zejian. The signaling pathways of disease resistance in Arabidopsis[J]. Acta Phytopathol Sin, 2003, 33(2):104-111. [19] LIU Sixin, ANDERSON J A. Marker assisted evaluation of Fusarium head blight resistant wheat germplasm[J]. Crop Sci, 2003, 43(3):760-766. [20] THOMMA B P H J, EGGERMONT K, BROEKAERT W F, et al. Disease development of several fungi on Arabidopsis can be reduced by treatment with methyl jasmonate[J]. Plant Physiol Biochem, 2000, 38(5):421-427. [21] BERROCAL-LOBO M, MOLINA A. Ethylene response factor 1 mediates Arabidopsis resistance to soilborne fungus Fusarium oxysporum[J]. Mol Plant-Micr Int, 2004, 17(7):763-770. [22] 高海波, 沈应柏.水杨酸甲酯、苯骈噻唑及茉莉酸甲酯对合作杨防御物质的影响[J].西北林学院学报, 2007, 22(6):32-35. GAO Haibo, SHEN Yingbai. Effects of BTH, MeSA and MeJA on defensive components of Populus simonii×P. pyramidalis[J]. J Northwest For Univ, 2007, 22(6):32-35. [23] 张可文, 安钰, 胡增辉, 等.脂氧合酶、脱落酸与茉莉酸在合作杨损伤信号传递中的相互关系[J].林业科学研究, 2005, 18(3):300-304. ZHANG Kewen, AN Yu, HU Zenghui, et al. Relationship between lipoxygenase and ABA and JA in wounded signal transduction of heathy populus seedlings[J]. For Res, 2005, 18(3):300-304. [24] CHENG Qiang, ZHANG Bo, ZHUGE Qiang, et al. Expression profiles of two novel lipoxygenase genes in Populus deltoides[J]. Plant Sci, 2006, 170(6):1027-1035. [25] HOOYKAAS P J J, BEIJERSBERGEN. A G M. The virulence system of Agrobacterium tumefaciens[J]. Annu Rev Phytopathol, 1994, 32(1):157-179. [26] PRADEL K S, ULLRICH C I, CRUZ S S, et al. Symplastic continuity in Agrobacterium tumefaciens-induced tumors[J]. J Exp Bot, 1999, 50(331):183-192. [27] DITT R F, NESTER E W, COMAI L. Plant gene expression response to Agrobacterium tumefaciens[J]. Proc Nat Acad Sci USA, 2001, 98(19):10954-10959. [28] VEENA, JIANG Hongmei, DOERGE R W, et al. Transfer of T-DNA and Vir proteins to plant cells by Agrobacterium tumefaciens induces expression of host genes involved in mediating transformation and suppresses host defense gene expression[J]. Plant J, 2003, 35(2):219-236. [29] LORENZO O, PIQUERAS R, SáNCHEZ-SERRANO J J, et al. Ethylene-response-factor 1 integrates signals from ethylene and jasmonate pathways in plant defense[J]. Plant Cell, 2003, 15(1):165-178. [30] GU Yongqiang, WILDERMUTH M C, CHAKRAVARTHY S, et al. Tomato transcription factors Pti4, Pti5, and Pti6 activate defense responses when expressed in Arabidopsis[J]. Plant Cell, 2002, 14(4):817-831. -

-
链接本文:
https://zlxb.zafu.edu.cn/article/doi/10.11833/j.issn.2095-0756.2014.05.009







 下载:
下载: